Somatic mutation rates scale with lifespan across mammals
Surveying great inventors and businesses
Axial: https://linktr.ee/axialxyz
Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company. We are excited to be in business with you — email us at info@axialvc.com
This study provides insights into the landscape of somatic mutations across 16 mammalian species, revealing a striking inverse relationship between somatic mutation rates and species lifespan. Using laser capture microdissection and whole-genome sequencing of 208 individual colorectal crypts from 56 individuals across the 16 mammalian species. Focusing on colorectal crypts with previous work in humans showing that they become clonally derived from a single stem cell and accumulate mutations linearly with age.
By carefully controlling for factors like tissue degradation time and using robust bioinformatic pipelines to call variants despite variable genome assembly quality, the authors generated high-quality somatic mutation data across a diverse set of species. The use of species with chromosome-level genome assemblies also enabled analysis of large-scale copy number changes.
Despite the diversity of species examined, the authors identified remarkable similarities in their mutational spectra and the major mutational processes contributing to somatic mutations. Three main mutational signatures – SBS1 (deamination of 5-methylcytosine), SBSB (resembling human SBS5), and SBSC (oxidative damage, like SBS18) – dominated across species, although their relative contributions varied.
This conservation of major mutational processes suggests that the core mechanisms of endogenous mutagenesis are largely preserved across mammals, likely reflecting conserved DNA metabolic pathways. The subtle variations observed, such as in the SBSB signature profile, hint at some species-specific differences that could be further explored.
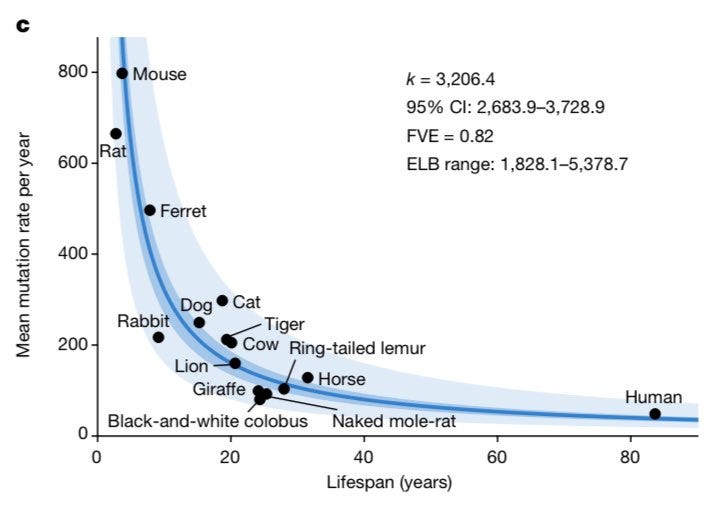
The most striking finding is the strong inverse scaling of somatic mutation rates with species lifespan. Across the 15 species with age data, substitution rates per genome per year ranged widely, from 47 in humans to 796 in mice. However, these rates exhibited a tight log-linear anticorrelation with species lifespan, with a slope close to -1.
This inverse proportionality suggests a simple model where the total somatic mutation burden (or end-of-lifespan burden, ELB) per cell is relatively constant across species, varying only by about 3-fold despite a 30-fold range in lifespan and 40,000-fold range in body mass. The authors' rigorous statistical analyses, employing linear mixed-effects models and accounting for phylogenetic relationships, strongly support this inverse scaling between mutation rate and lifespan, rather than with other life-history traits like body mass or metabolic rate.
The observed anticorrelation directly aligns with the long-standing "somatic mutation theory of ageing," which posits that accumulating somatic mutations contribute to the aging process and are therefore evolutionarily constrained, particularly in longer-lived species. As the authors discuss, this interpretation is consistent with evolutionary models predicting selection against life-threatening damage up to a species' typical lifespan, but weaker selection beyond that point.
While the study cannot prove causation, the remarkable quantitative fit to theoretical predictions lends strong support to the idea that somatic mutations play a role in aging, and that mutation rates have been modulated by natural selection across species. Alternative explanations, such as variable cell division rates or indirect effects of germline mutation rates, seem less plausible given the data.
Interestingly, the anticorrelation held not just for the overall mutation rate, but also for individual mutational signatures like SBS1 and SBSC thought to arise from distinct biological processes. This suggests that multiple DNA repair pathways may have been evolutionary tuned, rather than a single process driving the effect.
While the study focused primarily on the aging context, the authors also discussed the implications for the long-standing "Peto's paradox" in cancer evolution. The paradox notes the lack of correlation between body mass (and theoretical number of cellsat risk) and cancer incidence across species, despite the expectation from multi-stage cancer models.
The authors' finding that somatic mutation rates exhibit a much weaker correlation with body mass than with lifespan suggests that lowering mutation rates may not be the principal anti-cancer mechanism utilized during evolution of larger body plans. This implies that additional or alternative strategies, such as enhanced tumor suppression or elimination of mutant cell lineages, may have been pivotal adaptations permitting the evolution of larger bodies.
The apparent rarity of somatic copy number alterations and driver mutations in normal tissues appears conserved across mammals, consistent with prior observations in humans. Unlike humans, there was little evidence of mutational signatures like colibactin and APOBEC across the non-human species examined, suggesting differences in mutagenic exposures. While mitochondrial mutation rates were difficult to quantify precisely, the authors detected a substantial mitochondrial mutation burden (averaging 0.23 mutations per mtDNA copy) that also appeared to anticorrelate with lifespan. Curiously, despite their vastly different body sizes, some species like the giraffe and naked mole-rat exhibited very similar somatic mutation rates in line with their comparable lifespans, highlighting lifespan as the stronger predictor.
A key strength of this study is its pioneering cross-species comparative approach and robust statistical analyses on a expansive dataset. By examining somatic mutations in multiple individuals across diverse mammalian species, spanning orders of magnitude in body masses and lifespans, the authors could disentangle covarying life-history traits and unmask the predominant role of lifespan.
However, the study focused only on a single tissue type – the colorectal epithelium. While a reasonable choice given its clonal architecture and lower environmental exposure, extending such analyses to other tissues could prove informative. Second, the limited sample sizes for some species hindered detailed exploration of aspects like mutational signature variations and selection on coding mutations. Lastly, as the authors acknowledge, while the results associate somatic mutation rates with lifespan, conclusively demonstrating a causal role in aging will require further experimental evidence.
Ultimately, integrating such somatic mutation data with studies on aging phenotypes, stem cell dynamics and tissue dysfunction across species may help elucidate if and how somatic mutations causally contribute to aging processes. The authors demonstrated an inverse correlation between species lifespan and the rate of mutation accumulation between species. Notably, naked mole rats have an extremely low absolute number of mutations per genome compared to mice. This raises the question of whether they possess a more faithful DNA polymerase, but without mechanistic evidence, the accumulation of somatic mutations could be a consequence rather than a driver of aging.
While animals ultimately accumulate around 4,000 mutations per genome, over 90% of our genome does not code for proteins. Therefore, we might say that only 400 of these mutations target genes. However, out of the 20,000 genes in our genome, one cell type uses less than half of them. Thus, we are left with only around 200 plausibly detrimental mutations. Furthermore, not all mutations are missense, reducing the number further. Additionally, our genome is diploid, and for the vast majority of genes, one functional copy is sufficient. The probability of hitting one of the rare genes for which the loss of a single copy results in a strong phenotype, or hitting both copies of one gene among these 150 mutations, is low.
Moreover, many proteins have isoforms with partially or even fully overlapping functions. It's also worth noting that in this study, the authors used intestinal cells, a tissue known for one of the highest mutation rates in the organism. While the author expresses uncertainty and acknowledges their lack of expertise in genomics, they raise valid points. They mention the strong correlation between an increased number of mutations in a cell and augmentation of transcriptional noise. They also argue that untranslated regions of the genome might be less susceptible to DNA damage due to passive protection by histones, potentially targeting actively translated (and hence pivotal for cell function) regions of the genome with higher probability. The same could apply to enhancer regions, although their length is unknown. Finally, the author notes that the patterns recognized by transcription factors are very short. Such an integrative understanding could have profound implications for interventions aimed at delaying aging and age-related diseases.