Axial: https://linktr.ee/axialxyz
Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company. We are excited to be in business with you — email us at info@axialvc.com
This paper describes the design, synthesis, and validation of a large (3.7 million member) DNA-encoded library (DEL) called DOSEDO that incorporates diverse skeleton architectures derived from diversity-oriented synthesis (DOS). The authors created a DEL that captures more three-dimensional chemical space and structural diversity compared to traditional DEL synthesis approaches based on varying appendages on a common central scaffold.
Despite their power, most reported DELs have been limited by the need for DNA-compatible reaction conditions and common central scaffolds varied only by appendages. Using a branching synthetic approach where multiple diverse skeletons with varied exit vectors were elaborated with common appendage sets. This allowed greater 3D diversity compared to traditional DEL syntheses using a single central scaffold. 61 skeletons bearing secondary amines and aryl halides were selected, along with 295 amine capping agents and 206 boronic acids/esters for diversification.
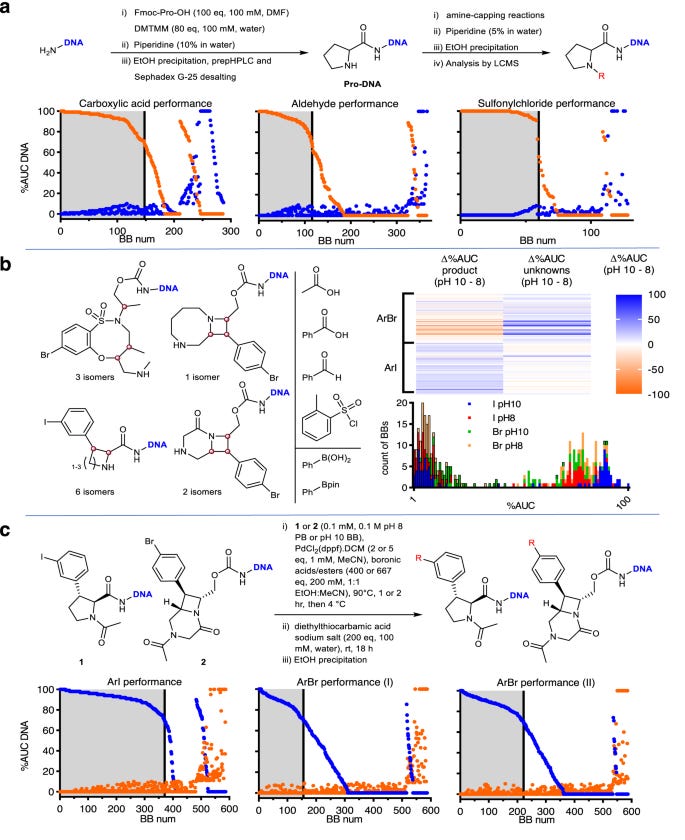
Significant synthetic method development is described. This included optimizing DNA conjugation of skeletons via carboxylates or hydroxyls, sulfonamide deprotection, acylation to minimize acetate byproducts, reductive amination, sulfonylation, and Suzuki coupling conditions using different catalysts, solvents, and reaction times. Validation of individual building blocks, skeletons, and reactions was thoroughly conducted.
Analysis of the validation data provided insightful observations. While amine capping reactions gave good overall coverage of chemical space, Suzuki couplings tended to work better for building blocks nearer the core of chemical space based on fingerprint analysis, with more outlier structures giving poorer results, especially for heterocycles and vinyl boronates. For different skeleton classes, some systematic effects were noted, like trans-piperidines performing poorly in reductive aminations. The authors carefully considered potential reasons for newly observed substructure bias beyond just steric and electronic effects.
Library synthesis started from the optimized conditions, generating separate sub-libraries for aryl iodide and bromide pools that were later combined. Barcoding and amplification for sequencing validation is described. Input sequencing revealed the iodide pool was over-represented, allowing a corrected pooling ratio for the final DOSEDO v2 library. Only 0.03% of expected sequences were not observed, some showing potential scaffold or reaction step bias worth future study.
Three protein targets were screened against DOSEDO: carbonic anhydrase IX (CAIX), isocitrate dehydrogenase (IDH1) R132H mutant, and ubiquitin-specific protease 7 (USP7). CAIX showed the expected enrichment for sulfonamides but also false negatives suggesting incomplete synthesis of some compounds. For IDH1, 9/25 resynthesized hits bound by SPR, with a sultam series showing good potency and enzymatic inhibition consistent with other inhibitors. For USP7, an enriched compound bound the GST tag unexpectedly, illustrating the range of possible screening outcomes.
Overall, this work significantly advances DEL technology by incorporating DOS skeletons to increase 3D diversity. Key strengths include the extensive synthetic methods developed, thorough validation of individual library components, and demonstration of protein binding hit identification. Potential limitations are the smaller size compared to some DELs, lack of optimization for the protein screens, and residual bias observed for some skeletons/reactions.
A few areas could potentially be expanded further:
1) More analysis correlating scaffold/reaction effects with calculated properties beyond just fingerprints. This could provide additional insight into synthetic biases.
2) More details on library production scale, final characterization methods, and storage/distribution plans for facilitating academic access as proposed.
3) Ideas for further enhancing scaffold/appendage diversity beyond the building blocks used here, or thoughts on ideal DEL size/complexity.
4) Discussion of the approach's potential limitations compared to conventional synthesis/screening, or opportunities for future integration of these methods.
5) Estimation of the work's overall cost and resource requirements as a potential barrier for some academic groups.