Axial - University of Chicago #1
Analysis of exciting University of Chicago life sciences inventors and their inventions
University of Chicago is where fun goes to die. Financed by John D. Rockefeller, Silas Cobb, and Marshall Field, the university was the epicenter for the Manhattan project. For life sciences, the university has churned out incredible companies maybe not at the Manhattan project-level yet but with a lot of promise. Actually, with the best so far likely ARCH Venture Partners, an investment firm not really a traditional company.
Kraig Lab
Studying how the brain heals itself.Recent
Demonstrating intranasal delivery of insulin-like growth factor-1 (IGF-1) and its benefits of reducing the underlying cause of migraines, slow depolarization in gray matter of the brain - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5993215/ - forming the basis of Seurat Therapeutics:
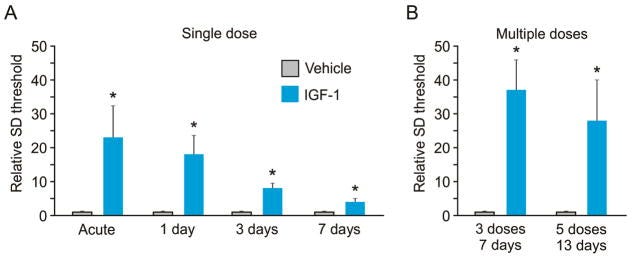
Past
Characterizing the role of exosomes derived from immune cells increased myelin levels and reduce neuronal stress - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4860060/ - very promising for companies like Mantra Bio and Codiak.
Studying IFNγ signaling and its role in migraines - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4589269/
Gajewski Lab
Studying T-lymphocytes and engineering them to target cancer.Recent
Overview on how the innate immune system particularly the STING pathway (cytosolic DNA sensing) can be engineered to treat cancer -https://onlinelibrary.wiley.com/doi/abs/10.1111/imr.12765
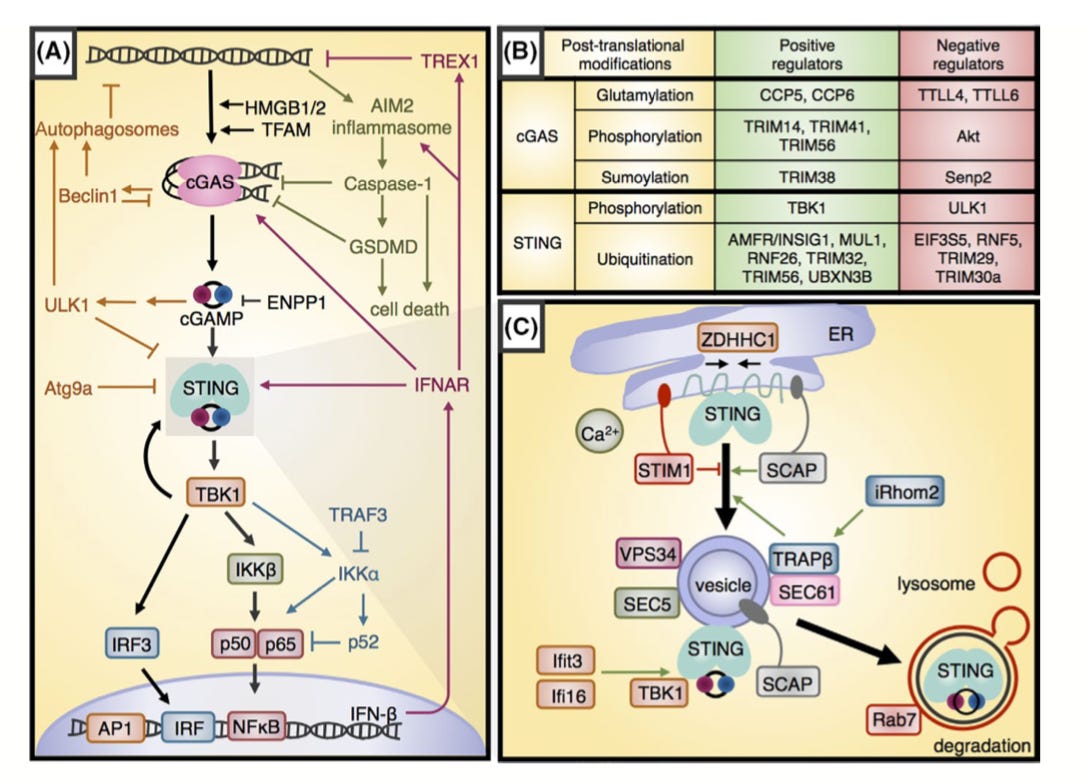
The Gajewski Lab is a pioneer in immunotherapy - just relatively ignored, which makes for a great opportunity; recently screened hundreds of neoantigens (specific targets for cancer cells) for not only binding but thermal stability relying on differential scanning fluorimetry -https://cancerimmunolres.aacrjournals.org/content/7/1/50.long - which is useful to infer binding affinities and cell surface expressivity; now acting as the core for Pyxis Oncology:
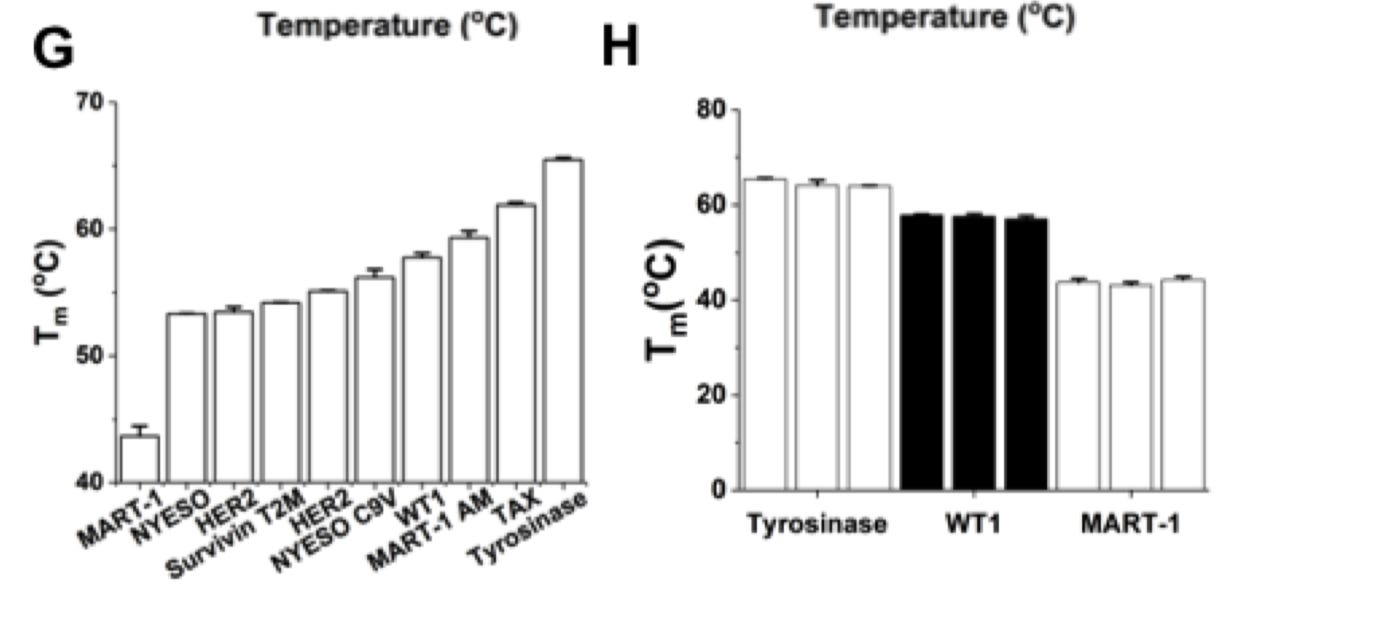
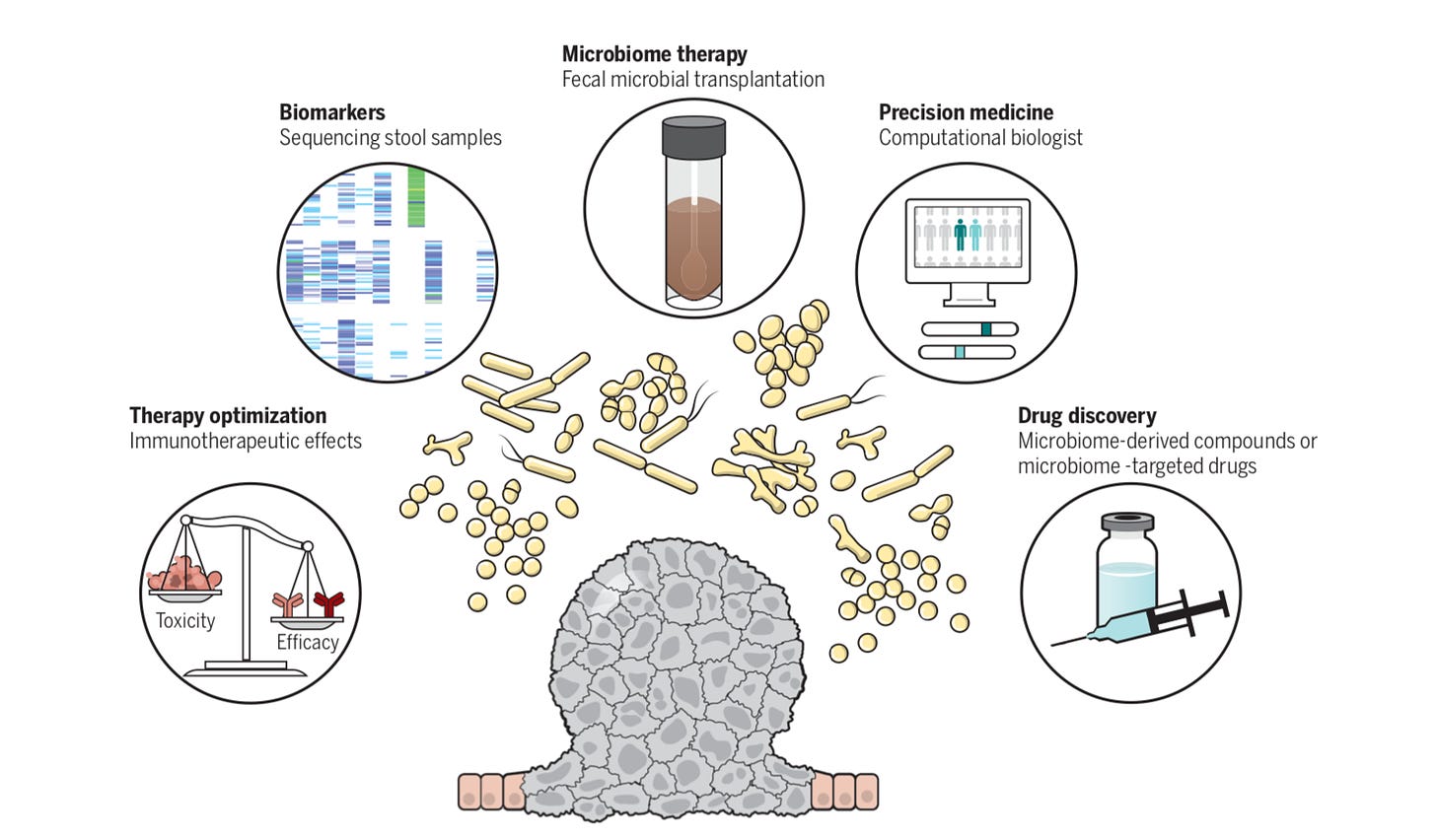
Overview on the role of the microbiome for immunotherapies (hopefully Persephone can make it work) -https://science.sciencemag.org/content/359/6382/1366.long
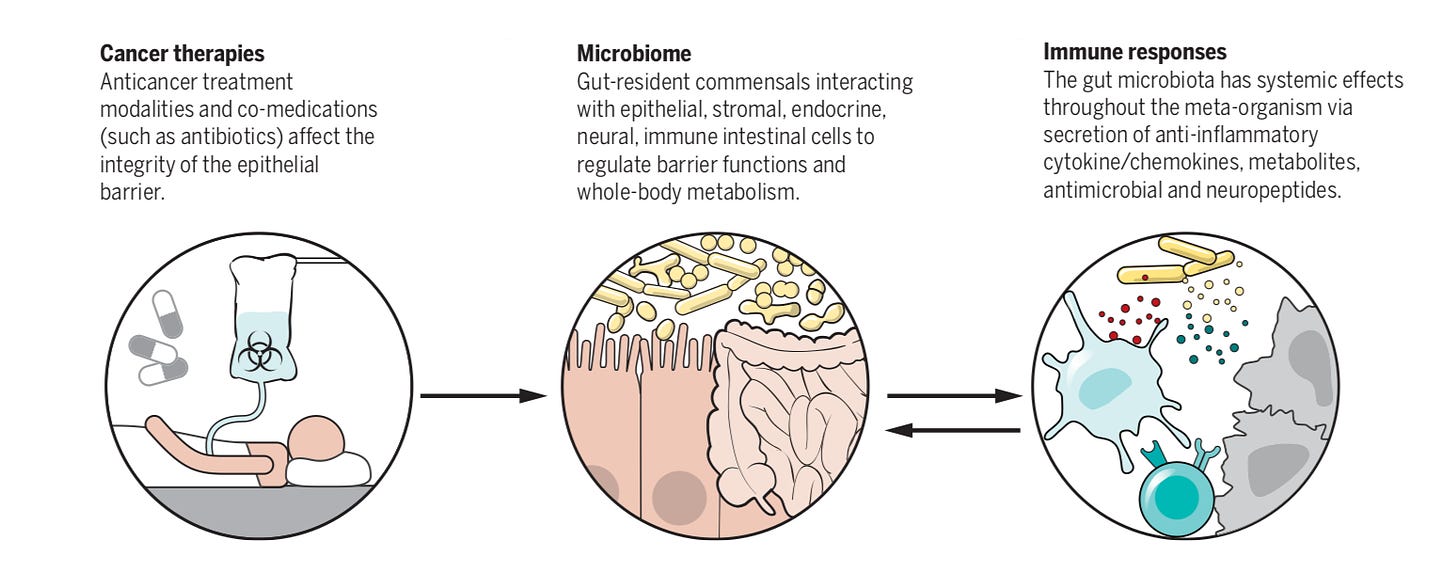
Tay Lab
Modeling living systems to predict their behavior.Recent
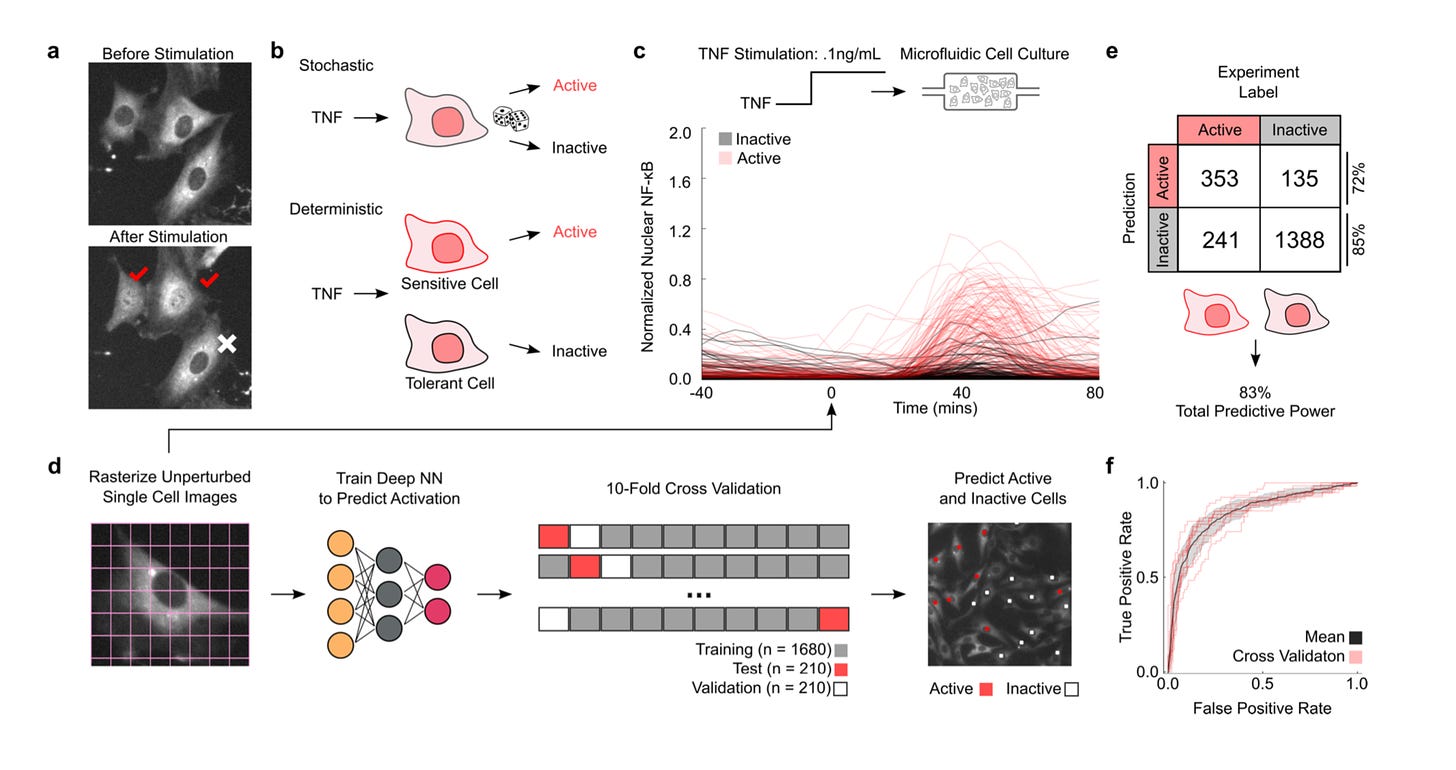
Doing really exciting work in biological engieering - already helped formed BiomeSense, but the Tay Lab has more exciting inventions within its keep. Invention a computer vision method to measure NF-κB signalling and determine a given cell or classes ability to respond to TNF - https://www.biorxiv.org/content/10.1101/687848v1 - establishing a method that can be used for general drug screening:
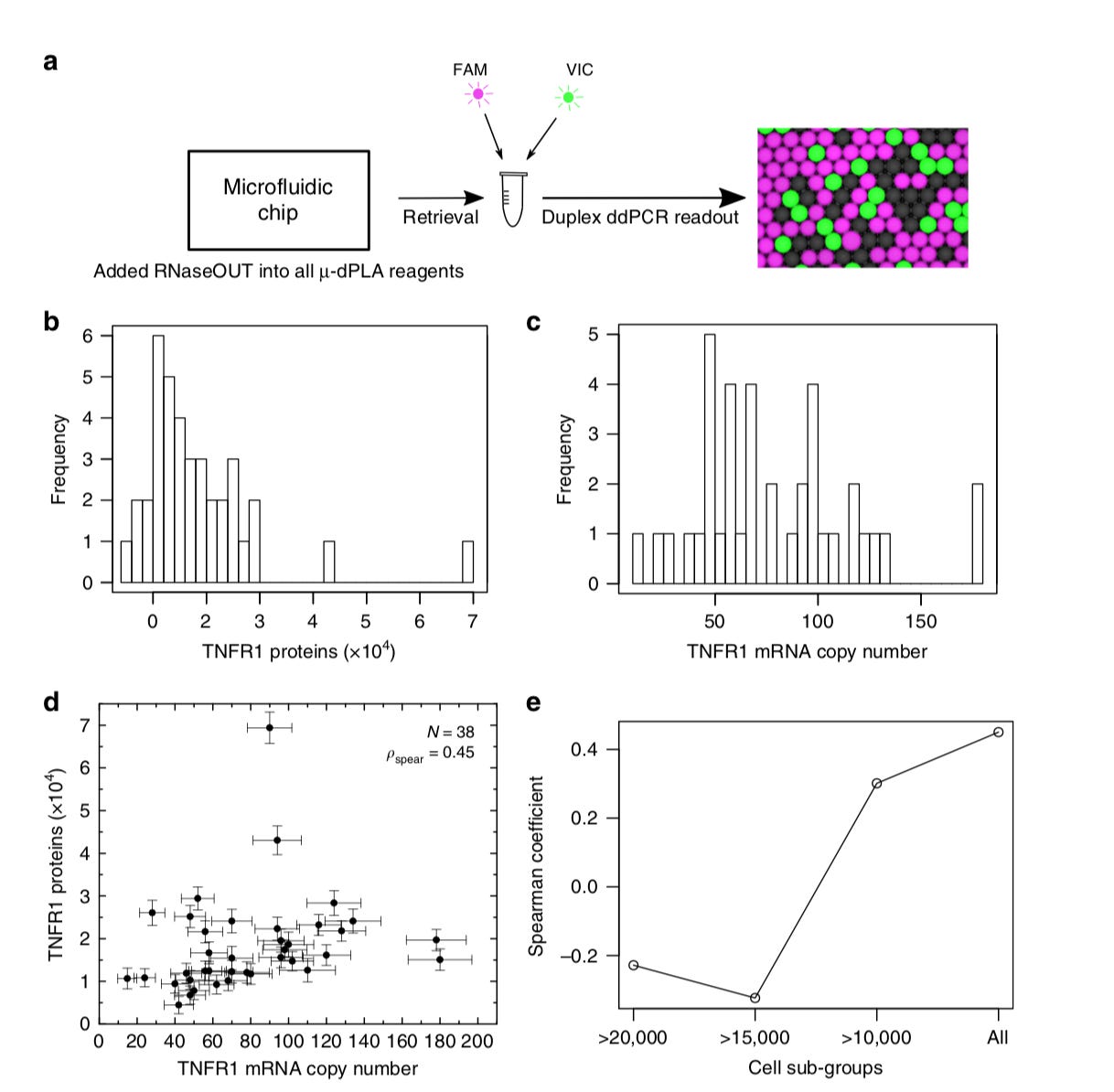
Another exciting piece of research using microfluidics to measure single-cell proteomes and mRNA populations -http://taylab.uchicago.edu/uploads/9/1/8/0/91804060/2019-JL-NatComm.pdf
Hassan Lab
Understanding how waste in the body is transported and regulated.Recent
A young lab with a lot of potential to look and potentially disease from a new angle - physiological waste. Whose work is supporting Oxalo Therapeutics to treat recurring kidney stones. Studying how pro-inflammatory cytokines drive oxalate secretion (kidney stones are made up of calcium oxalate) -https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5963707/
Past
Screening for factor that drive oxalate transport -https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5328155/
Drummond Lab
Engineering translation at scale.Recent
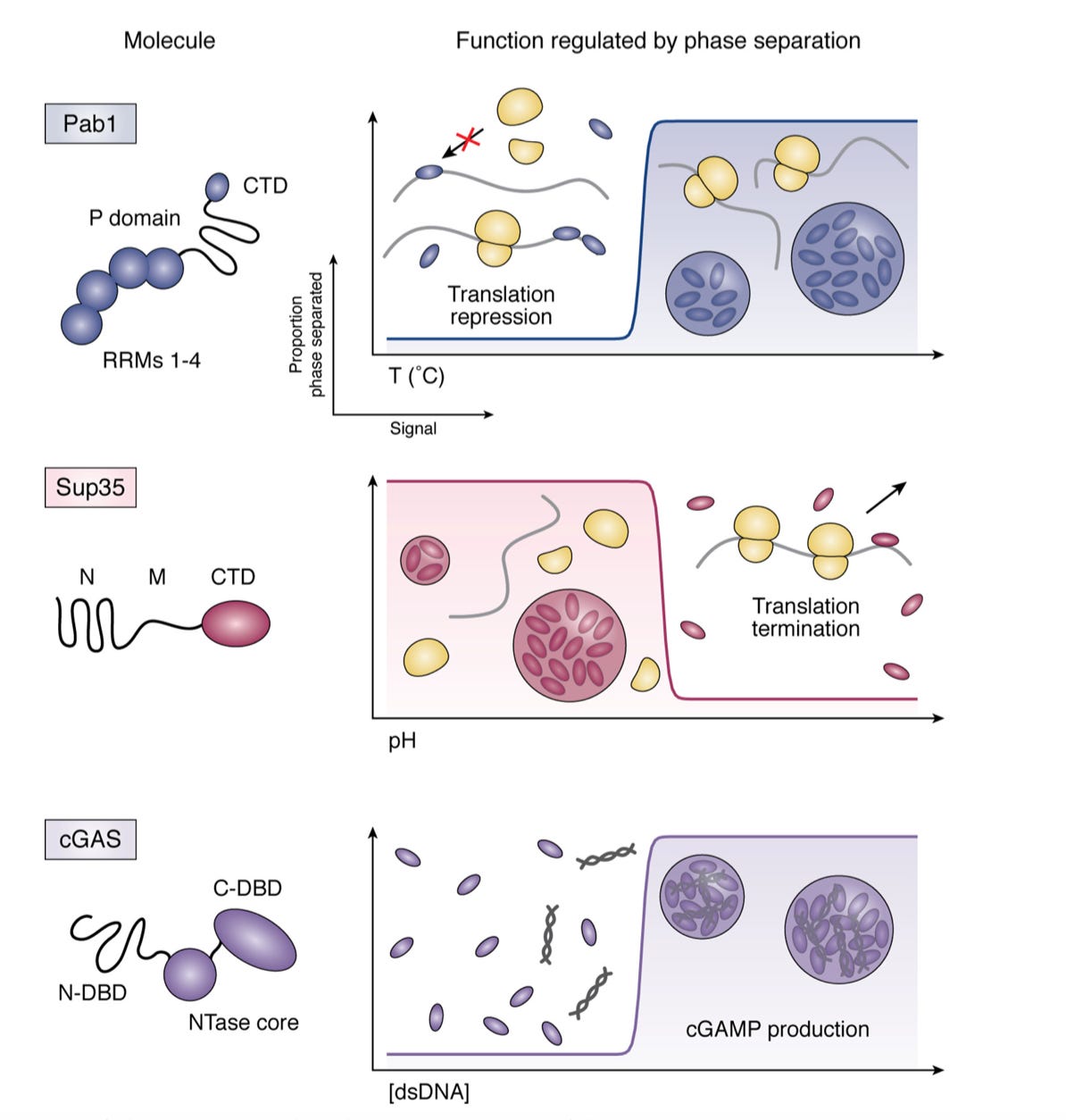
A fascinating review on understanding and potentially engineering biological phase separation . Increasingly, especially for transcription, phases and condensates within cells are found to have importance in imbuing selectively and sense environmental changes. Given the Drummond Lab’s expertise considerable depth is given in the role of phase separation for translation -http://www.jbc.org/content/294/18/7151.full.pdf
Past
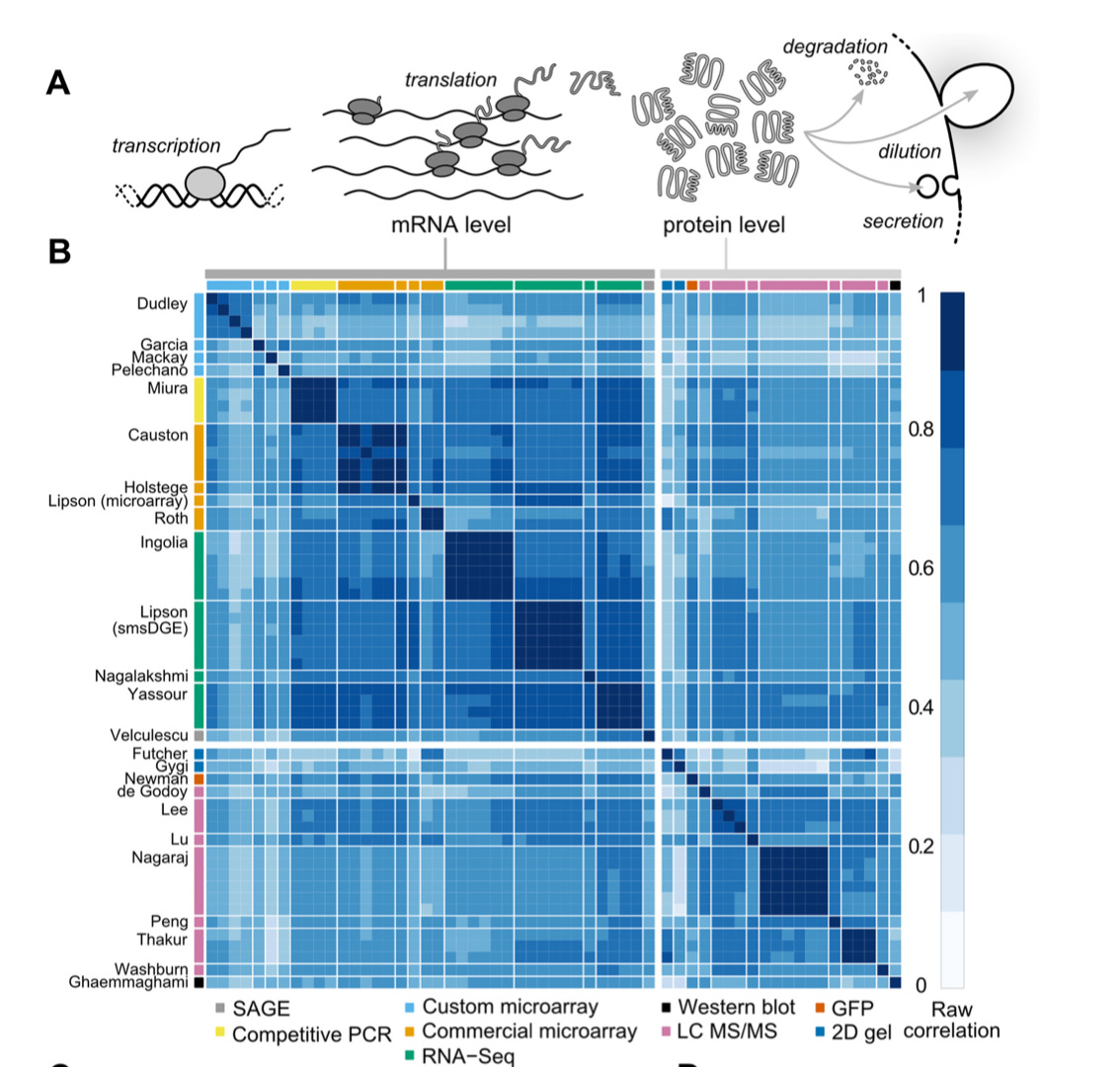
In a tour de force, analyzing comprehensive mRNA and protein levels in yeast across 24 studies - http://www.plosgenetics.org/article/fetchObject.action?uri=info:doi/10.1371/journal.pgen.1005206&representation=PDF - discovering that mRNA levels determine the majority of protein levels (albeit at equilibrium) instead of the common-held belief at the time that mRNA only determines a little over a third. This revision showed that modulating mRNA, the bottleneck for proteins, is a worthwhile pursuit for drug development:
Nagler Lab
Characterizing self and non-self within the microbiome.Recent
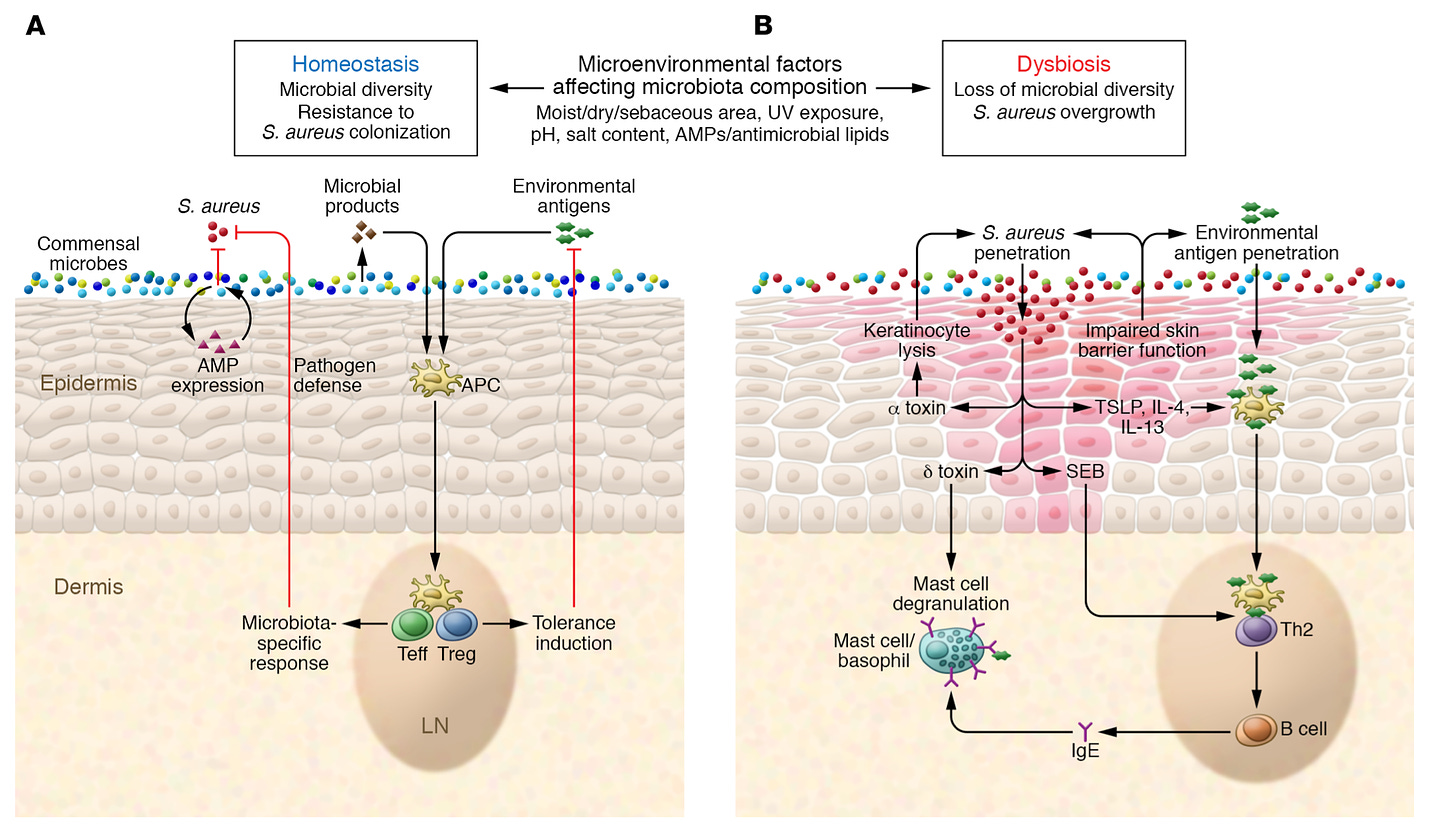
A legendary lab, recently putting out a review on the role of the human microbiota for allergies - https://cpb-us-w2.wpmucdn.com/voices.uchicago.edu/dist/e/1480/files/2019/07/Kemter-Nagler-2019-Influences-on-allergic-mechanisms-through-gut-lung-and-skin-microbiome-exposures.pdf
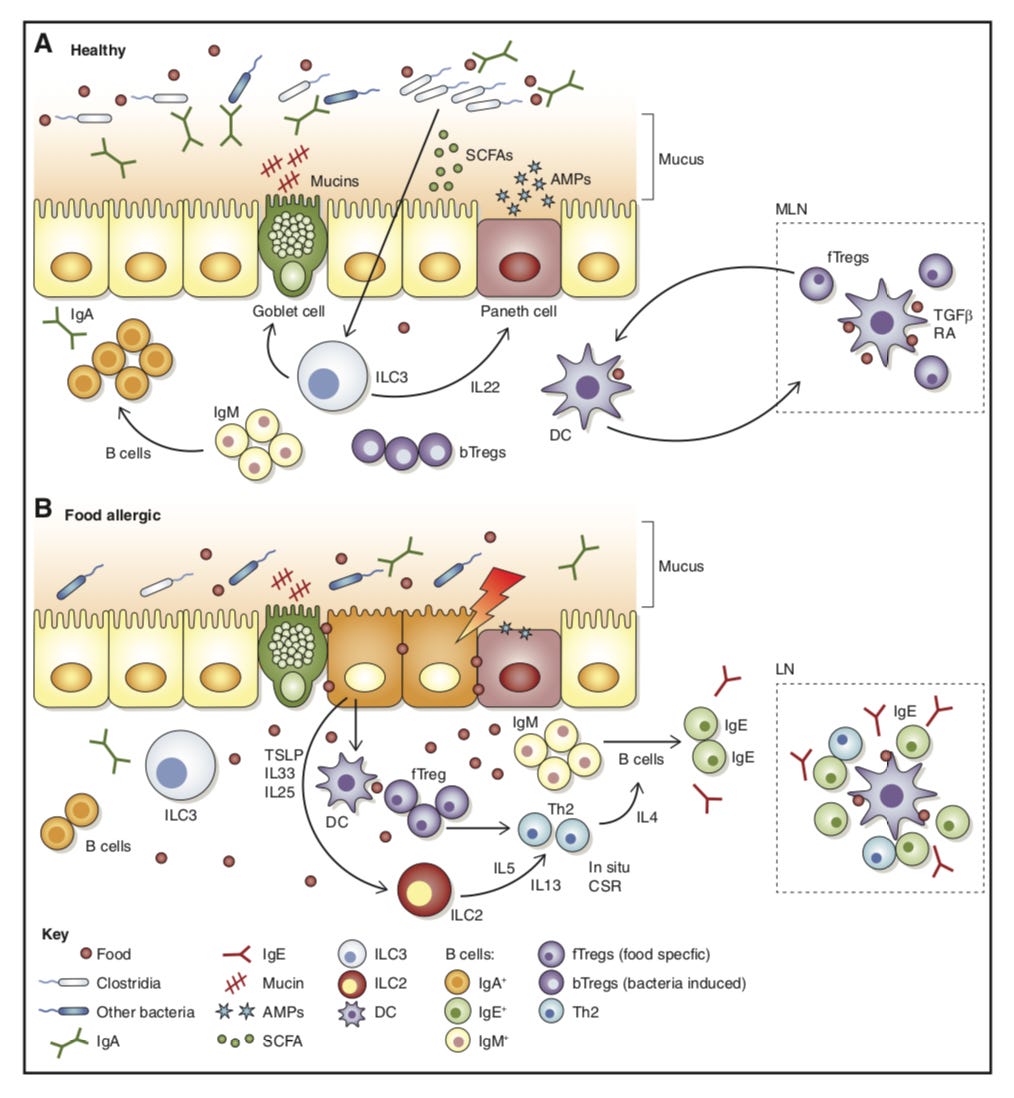
In mice, discovering that Anaerostipes caccae protected against an allergic response to food - https://cpb-us-w2.wpmucdn.com/voices.uchicago.edu/dist/e/1480/files/2019/07/Feehleyetal.pdf - another brick in the wall toward definitively treating allergies and even autoimmunity in general via food; hopefully Ukko can bring something useful to market.
Past
Showed that Clostridia-containing microbiota protected against allegies in mice - http://clostrabio.com/uploads/3/6/1/9/36196116/stefka_pnas_commensalbacteria_2014.pdf - forming the basis of ClostraBio.
A valuable review on the role of the microbiome in allergies -http://clostrabio.com/uploads/3/6/1/9/36196116/wesemann_immunity_microbiomecontext_2016.pdf
Adams Lab
Understanding how the human immune system determines self from non-self.Past
Implementing an unbiased T-cell receptor (TCR) screen to understand how CD1 presents Mycobacterium tuberculosis antigen selectively - https://www.jimmunol.org/content/196/4/1933 - incredibly important to understand self/non-self interactions in the CD1 system.


