Axial - Inventors #25
Surveying great inventors and businesses
Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company . We are excited to be in business with you - email us at info@axialvc.com
Inventors #25
A set of ideas and observations on inventions and discoveries in life sciences.
Immunology
The immune system and everything in it.
Succination inactivates gasdermin D and blocks pyroptosis - https://science.sciencemag.org/content/369/6511/1633 - the Fitzgerald Lab at UMass Medical School discovered that dimethyl fumarate (DMF) succinates gasdermin D (GD) explaining the drug’s mechanism treating multiple sclerosis as well as opening up another avenue to selectively target GD:
Gasdermin D is an exciting drug target involved in multiple inflammatory cell death pathways. In general, the gasdermin class is found in the GI, skin, and mucosal and immuno-inflammatory cells.
GD cleavage induces pyroptosis (N-terminal domain responsible for initiation and the C-terminus inhibits the N-terminus). Recent structural work has been transformational for the field - https://www.sciencedirect.com/science/article/pii/S1074761319301979#
In 2015, work out of Genentech established the role of gasdermin D in pyroptosis for one of the biggest breakthroughs in inflammation over the last few decades - https://www.nature.com/articles/nature15541 In short, when pyroptosis is activated, gasdermin D forms lytic pores on the cell membrane to mediate the release of cytokines and other inflammatory signals. So this immediately made GD an interesting drug target for diseases ranging from lupus to NASH and multiple sclerosis.
The Fitzgerald Lab discovered the role of dimethyl fumarate (DMF) and endogenous fumarate disrupts the process of GD forming pores through succination (a PTM on cysteines)
Through screening, the group discovered that DMF inhibited the activation and cleavage of GD’s N-terminal by Caspase-11. Then they discovered this MoA is mediated by succination and identified gasdermin D as a target of the fumarate class of molecules.
This is an important paper to understand how to successfully drug GD since it’s been pretty tough so far
The general idea is to develop a GD inhibitor for inflammatory diseases where NLRP3 or inflammasome inhibition is not sufficient
Biochemistry and structural biology
The granddaddy of them all.
Protein design-scapes generated by microfluidic DNA assembly elucidate domain coupling in the bacterial histidine kinase CpxA - https://www.pnas.org/content/118/12/e2017719118 - out of the Abate Lab at UCSF, the group invented a microfluidic-base method to construct libraries of multi-domain proteins:
Generating comprehensive libraries of proteins is very useful since it is difficult to predict which protein sequences are functional and which ones are not. Instead of predicting sequence-function relationships, just printing out each protein variant and testing them sidesteps this problem.
The group designed and implemented a multi-valve microfluidic device. I haven’t made a microfluidic device in ~6 years so I am a bit rusty on the paper’s process, but in short, the device can mix-and-match 382 pL droplets from 38 inputs (10 ms response times) to generate ~10K protein variants in about 2 hours.
So the paper used this device to generate a combinatorial library of histidine kinase, CpxA to find variants with higher kinase activity
Scanning across several CpxA domains discovered protein variants with higher catalytic activity from differential S-helix length (involved in regulation with the TM domain)
This study established a new method to generate protein libraries. Next steps are scaling but there are obvious applications in the design of multi-domain therapeutics. Great work.
Neuroscience
Roughly 20 years behind but set up to transform the concept of human.
Deep sequencing of sncRNAs reveals hallmarks and regulatory modules of the transcriptome during Parkinson’s disease progression - https://www.nature.com/articles/s43587-021-00042-6 - the Keller Lab at Saarland University used sequencing to identify blood circulating non-coding RNAs that have diagnostic potential for Parkinson’s disease:
The work was premised on past observations that microRNAs (miRNAs) and other non-coding RNAs (ncRNA) have aberrant expression in Parkinson’s (PD) and other neurodegenerative diseases
The group relied on sequencing data on over 5K samples across 1,614 patients from the PPMI and NCER-PD
PPMI was done to generate a public, high-quality datasets from RNA from whole-blood of PD patients across different stages of the disease
The group’s analysis of this dataset discovered that miRNA expression profiles can be used to classify patients with idiopathic versus genetic Parkinson’s
This analysis in conjunction with a literature search of miRNAs annotated as PD markers discovered that miR-15b-5p (in serum) downregulation is a possible predictive market of Parkinson’s along with miR-185-5p
This is really early analysis that still needs to be controlled for things like gender, age, and other patient attributes, but this work is a good first step toward identifying biomarkers for easier detection of neurodegeneration
Cell biology
Cell structure and function.
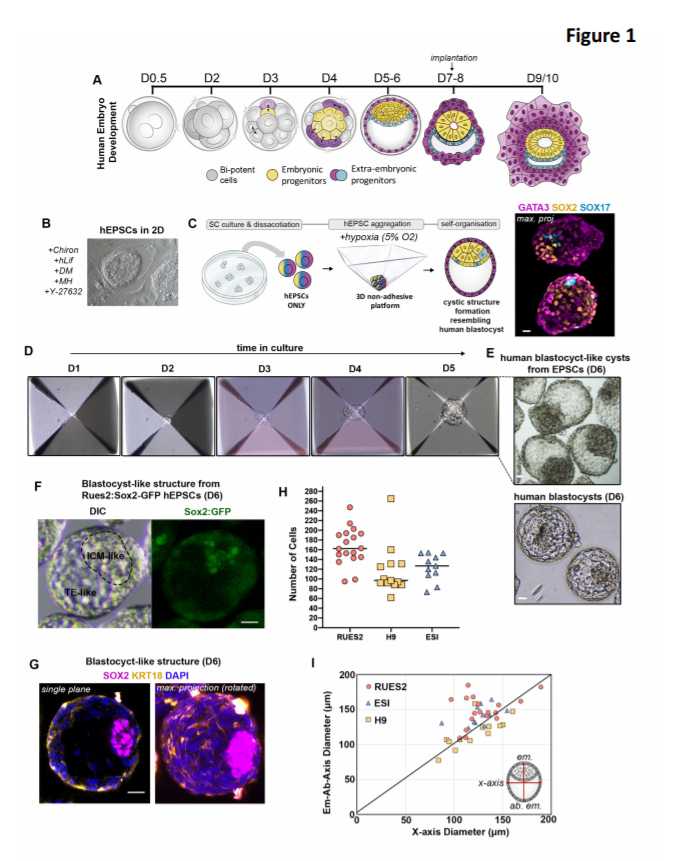
Reconstructing human early embryogenesis in vitro with pluripotent stem cells - https://www.biorxiv.org/content/10.1101/2021.03.12.435175v1 - the Zernicka-Goetz Lab at the University of Cambridge and Caltech used human pluripotent stem cells (hPSC) to reconstitute early human embryogenesis:
After conducting a systematic screen (3D culture system) of growth factors, cytokines, and small molecules, the group discovered a series of factors that mimic the environment of the early-stage embryo to guide PSCs to gain the physical features of human blastocysts
A combination of “BMP4 (20ng/ml), WNT antagonist CHIR99021 (2uM), FGF2 (40ng/ml) and ROCK inhibitor Y-27632 (5uM) during the first 48h of 3D culture enhanced cell survival and promoted formation of compact cellular aggregates”
To figure out if their systems recapitulates early embryogenesis, the paper performed qRT-PCR before and after aggregate formation for biomarkers indicative of blastomeres - ZSCAN4, THAP11 and NR5A210,26,31, which were expressed 2x to 8x higher before aggregate formation and were downregulated afterwards
This paper does a nice job at building a more accurate model of human blastocysts that can be used to study the developmental trajectories of single-cells and the regulatory system of human embryogenesis
Genetics, genomics, and developmental biology
Heredity and variation.
Programmed ER fragmentation drives selective ER inheritance and degradation in budding yeast meiosis - https://www.biorxiv.org/content/10.1101/2021.02.12.430990v1 - the Brar Lab at UC Berkeley did some incredible work in budding yeast to connect the morphology of the endoplasmic reticulum (ER) to its inheritance or selective degradation:
The paper sought out to understand what happens to the ER during meiosis?
Time-lapse imaging uncovered that parts of the cortical ER, the largest subdomain in yeast that is associated with the plasma membrane (PM), un-tethers from the PM during meiosis (anaphase II) based on the expression of a series of genes (Ndt80, Lnp1, among others). Parts of the ER were inherited across gametes and re-tethered to the PM.
To study how these untethered ER components are degraded, the group designed an elegant experiment to measure ER autophagy. Due to GFP being resistant to ER-resident proteases, ER-related proteins were tagged with GFP and measured GFP levels over meiosis. This work discovered that Atg40 (autophagy receptor) was necessary to degrade un-tethered ER.
Outstanding questions remain around selectivity of this ER quality control process, but the work is an outstanding step forward to link a biophysical event (ER tethering to PM) to inheritance