Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company . We are excited to be in business with you - email us at info@axialvc.com
Inventors #21
A set of ideas and observations on inventions and discoveries in life sciences.
Immunology
The immune system and everything in it.
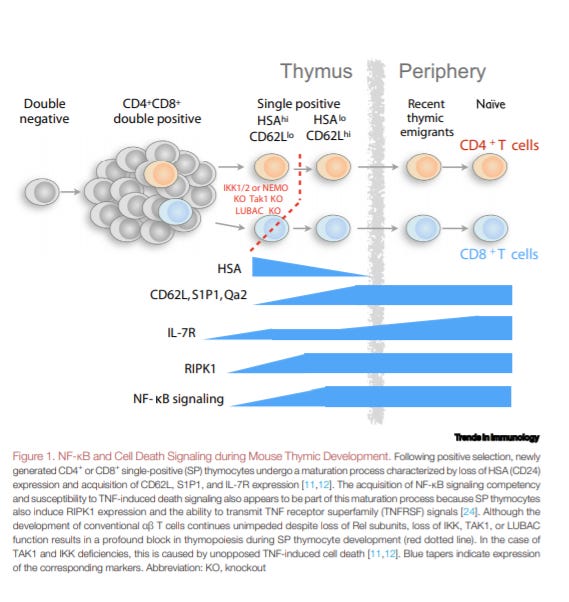
NF-κB and Extrinsic Cell Death Pathways – Entwined Do-or-Die Decisions for T cells - https://www.cell.com/trends/immunology/fulltext/S1471-4906(20)30257-X#%20 - a review from the Seddon Lab at UCL does a great job at characterizing the role of the NF-κB pathway in T-cell death:
NF-κB signaling is both pro- anti-apoptotic (i.e. cell death). In particular, the pathway is important for T-cell tolerance.
NF-κB family consists of five transcription factors: NF-κB1/p50, NF-κB2/p52, RelA/p65, RelB and c-Rel that homodimerize or heterodimerize to 15 NF-κB complexes. Various signals from cytokines to stress can activate various parts of this pathway. In terms of drug development, many different drugs indirectly target the pathway; the issue historically has been pleiotropy, which is a problem for all transcription factors. There is a need to connect a signal to a particular NF-κB model to a phenotype.
So the key observation the review made is that two kinases, IKK and TAK1, that regulate NF-κB have also been recently connected to cell death along with 3 downstream gene targets: A20, cIAP1, and cIAP2
Another interesting avenue of research is the role of NF-κB in non-effector T-cells in particular memory cells. Mice knockouts of NF-κB fail to develop memory T-cells.
Moreover, γδ T-cells require expression of CD27, which is strongly linked to NF-κB signaling
This is all done in mice - do the genetics match up to human immunology? The idea is to augment NF-κB signaling to decrease T-cell activity in autoimmunity or turn in up in an oncology setting.
The authors pose two key questions in the field: “Does NF-κB signaling contribute to long-term homeostasis of memory T cell populations?” and “How do NF-κB and cell death pathways combine to control the development and homeostasis of nonconventional T cells such as γδ, iNKT, MAIT, and IEL T cells?”
Biochemistry and structural biology
The granddaddy of them all.
Proteomic analysis of young and old mouse hematopoietic stem cells and their progenitors reveals post-transcriptional regulation in stem cells - https://elifesciences.org/articles/62210 - the Weissman Lab at Stanford did an elegant study using proteomics on old and young mouse hematopoietic stem cells (HSC):
This resource paper uses shotgun mass spectrometry on old and young mouse HSCs and progenitor cells across 12 different cell types to measure proteins that are measurable by mass spec (i.e. can be ionized): “young adult and old adult HSC (Lin- , cKit+ , Sca1+ , CD34- , CD150+ , Flt3- ), MPPa (Lin- , cKit+ , Sca1+ , CD34+ , CD150+ , Flt3- ), young adult and old adult MPPb (Lin- , cKit+ , Sca1+ , CD34+ , CD150- , Flt3- ), young adult and old adult MPPc (Lin- , cKit+ , Sca1+ , CD34+ , CD150- , Flt3+ ), young adult CLP (Lin, cKitlo, Sca1lo, Flt3+ , IL7Ra + ), young adult CMP (Lin- , cKit+ , Sca1lo/-, CD34med/hi , CD16/32-/lo), young adult MEP (Lin- , cKit+ , Sca1lo/-, CD34- , CD16/32-/lo, CD150+ ) and young adult GMP (Lin- , cKit+ , Sca1lo/-, CD34hi, CD16/32hi)”
The protocol relies on sorting cells by FACs and sample pooling to input a minimum of 50K cells for mass spec analysis
The dataset combined with RNA-seq data reveals a series of interesting observations on HSC development and self-renewal that require further study - detecting increased levels of CD150/Slamf1 and vWF in the old HSCs
Moreover, the paper observed higher levels of protein diversity in older HSCs versus younger ones. This might open up never avenues of research on stem maintenance. The author propose that stemness regulation shifts from transcription to translation over time
The paper also discovered that proteins associated with ubiquitin-driven degradation become less abundant in HSCs over time. As a result, other mechanisms of translation regulation might be more important such as miRNAs or lysosomal degradation
Neuroscience
Roughly 20 years behind but set up to transform the concept of human.
A non-hallucinogenic psychedelic analogue with therapeutic potential - https://www.nature.com/articles/s41586-020-3008-z - out of the Olson Lab at UC Davis, the group designed and synthesized a non-hallucinogenic psychedelic:
Ibogaine is a psychedelic that has been shown to potentially treat addiction. The group used Function-Oriented Synthesis - https://pubs.acs.org/doi/10.1021/ar700155p to identify the pharmacophore of ibogaine and develop a non-hallucinogenic version.
Beyond the small molecules they generate, the paper is a case study on the potential of developing psychedelic analogs for mental health
Key elements of ibogaine were discovered and were used to invent an analog called tabernanthalog
In rat models, the analog was discovered to promote neural plasticity and reduce addictive behaviors
In general, psychoplastogenic medicines have the potential to rewire neural circuitry and treat complex psychiatric disorders
Psychedelics like ketamine have shown efficacy signals in depression and other mental health disorders; however, the MoAs still have yet to be mapped out. Rational design of psychedelics are an important step to figure this out.
Next steps are figuring out dosing and connecting neural plasticity to endpoint surrogates
Cell biology
Cell structure and function.
Genetic interaction mapping informs integrative structure determination of protein complexes - https://science.sciencemag.org/content/370/6522/eaaz4910 - the Krogan Lab at UCSF use genetic interactions to infer the structure of protein complexes:
The method the group invented is a really neat approach to get to in vivo protein complex structures. This is pretty important given that for many structures of protein complexes that are discovered, it’s hard to know if they exist in the cell.
Building off their past work in genetic interaction mapping, the paper generated 479 mutants for the yeast histone H3-H4 complex, where 350 were not lethal and amenable to analysis, and crossed them against 1370 genetic deletions and hypomorphic alleles (important to map out PPIs with plausible proteins) to create a map of around 500K genetic interactions
The data from this screen was used to model the structure of the H3-H4 complex, which came out consistent with the known structure. The paper carried out this screen on 2 other complexes: subunits Rpb1-Rpb2 of yeast RNA polymerase II and subunits RpoB-RpoC of bacterial RNA polymerase.
This tool has the potential to become a critical complementary tool to determine the structure of protein complexes in cells in combination with X-ray crystallography and cryo-EM. In particular, this type of mapping could be useful to determine the structure of transient complexes or very large ones.
Next steps are to do this type of screening in mammalian cells and connect the structure of protein complexes to particular genetic mutations
Genetics, genomics, and developmental biology
Heredity and variation.
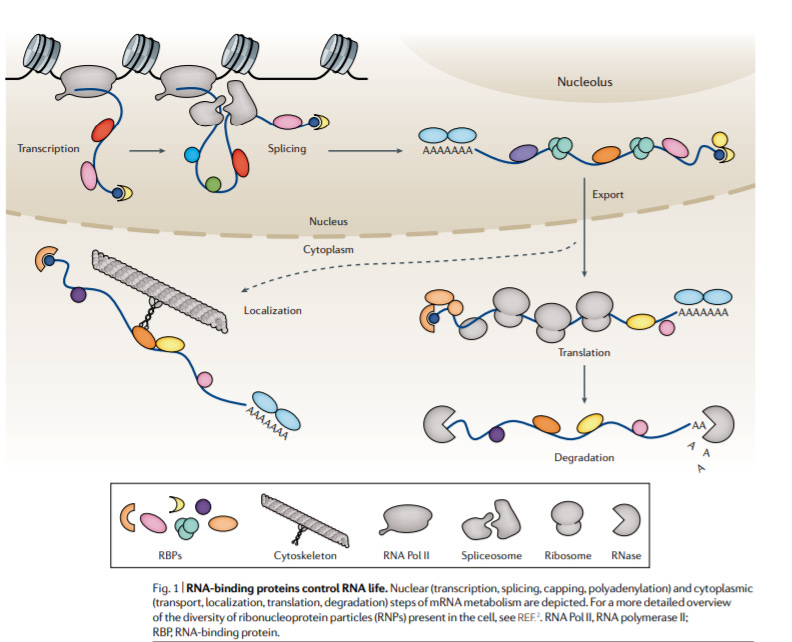
RNA-binding proteins in human genetic disease - https://www.nature.com/articles/s41576-020-00302-y - a useful review from the Hentze Lab at EMBL on the role of RNA-binding proteins (RBP) in disease:
There are thousands of known human RBPs with 1,054 of those mutated in disease, which represents over 20% of all disease-mutated proteins. Most studies rely on UV crosslinking to map out proteins that bind RNA, but there are probably low abundance RBPs that have been missed out on so far -
https://rbpbase.shiny.embl.de/
In particular, mutant RBPs are most associated with CNS diseases.
RBPs are important to form biomolecular condensates, regulate RNA translation and degradation, and metabolism.
A case study from the review is hydroxysteroid dehydrogenase (HSD17B10), which is mutated in mitochondrial cardiomyopathy/neuropathy syndrome. The mutation was discovered to decrease binding to mitochondrial pre-tRNAs.
With thousands of RBPs, there is a large opportunity to map out their roles in disease. It’s an open field where phenotypic screening, structural biology, and more are useful.
There is also an open question on intrinsically unstructured regions found within RBPs - how do these domains influence RNA binding?
Are they RBPs that can become attractive therapeutic targets?