Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company . We are excited to be in business with you - email us at info@axialvc.com
Inventors #16
A set of ideas and observations on inventions and discoveries in life sciences.
Immunology
The immune system and everything in it.
Modulation of PKM activity affects the differentiation of TH17 cells - https://stke.sciencemag.org/content/13/655/eaay9217 - the Gaultier Lab at the University of Virginia discovered small molecules that activate pyruvate kinase M2 block the differentiation of IL-17–producing Th17 cells, which are important for the pathology of multiple sclerosis (MS):
Pyruvate kinase (PK) is the rate-limiting for glycolysis to generate ATP
Immune cell activation is characterized by a metabolic shift from oxidative phosphorylation to glycolysis. This is known as the Warburg effect and is a hallmark in many cancers.
The goal of this work was to understand the therapeutic effects of targeting PK isoform M2 (expressed in immune cells) in MS
Several drug candidates in MS focus on shifting metabolism in immune cells with some activating PK. This paper challenges this approach by showing PK activation redirects inflammation from the spinal cord to the brain in a mouse model.
The key experiments showed that two PKM2 activators, TEPP-46 and DASA-58, suppressed differentiation of Th17 cells producing IL-17 but promoted those that produce GM-CSF, which is a hallmark of brain inflammation. Both cell types drive the pathology of MS; however, the PKM2 activators seem to suppress inflammation in the spinal cord (IL-17) and redirect it to the brain (GM-CSF).
Biochemistry and structural biology
The granddaddy of them all.
meCLICK-Seq, a Substrate-Hijacking and RNA Degradation Strategy for the Study of RNA Methylation - https://pubs.acs.org/doi/10.1021/acscentsci.0c01094 - out of the Bernardes Lab at the University of Cambridge, the group developed a new tool to monitor RNA methylation and degradation in cultured cells:
The tool, meCLICK-Seq (methylation CLICKdegradation Sequencing) use an S-adenosyl methionine (SAM) surrogate, SeAdoYn, to get RNA methyltransferases to use it as a cofactor and add a propargyl (an alkyne), instead of a methyl group, to RNA
After, the click degrader is added to the RNA through a Cu(I)-catalyzed azide−alkyne cycloaddition (CuAAC) reaction. The click degrader initiates RNA cleavages and leads to its degradation. The decrease in overall RNA methylation is used to monitor the modification for specific RNA transcripts.
The tool is unique because it relies on measuring depletion instead of enrichment of methylated RNA, does not rely on an antibody, and does not involved enzymatic processing - this enables meCLICK-Seq to measure a wide ranger, particularly rare species, of transcripts
Overall, meCLICK-seq is useful to measure RNA methylation as well as studying RNA methylases and mapping modification sites across both intronic and intergenic regions
Neuroscience
Roughly 20 years behind but set up to transform the concept of human.
A new neutrophil subset promotes CNS neuron survival and axon regeneration - https://www.nature.com/articles/s41590-020-00813-0 - the Segal Lab at The Ohio State University discovered a population of neutrophils with neuroprotective phenotypes in contrast to studies that show mature neutrophils are destructive in the CNS:
The group used a optic nerve crushing mouse model to study the role of immune cells in axon regeneration
Intraocular injection of zymosan, fungal extract, mitigates the effects of nerve crushing. This mechanism is mediated through the innate immune pattern recognition receptor, dectin-1, which is found on microglia and dendritic cells. With this observation, the group worked to understand the mechanism of regeneration through dectin-1.
Using this model and flow cytometry, a subset of immune cells (CD14+Ly6Glo) with features similar to neutrophils was discovered to have neuroprotective and axonogenic effects
This cell type was able to promote axon regeneration within the spinal cord, outside of the retina. This mechanism is thought to be done through the secretion of specific growth factors.
This is another step toward fully understanding axon regeneration and design new therapies to treat CNS injury
Cell biology
Cell structure and function.
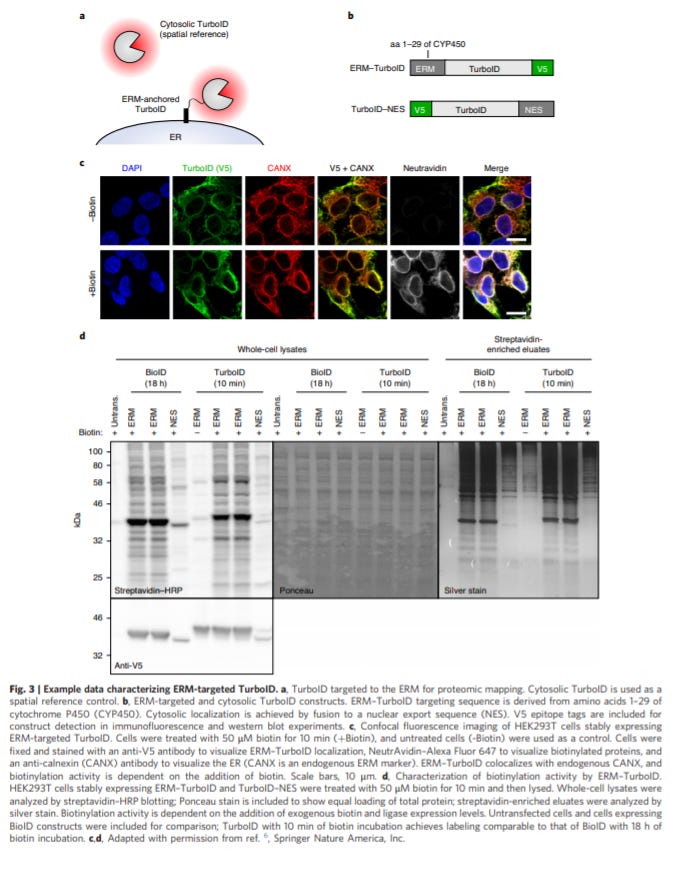
Proximity labeling in mammalian cells with TurboID and split-TurboID - https://www.nature.com/articles/s41596-020-0399-0 - from the Ting Lab at Stanford, the group invented a tool, TurboID, for more selective proteomic mapping with a focus on capturing rare protein-protein interactions:
The protocol lays the use of TurboID in cultured mammalian cells
The method uses an biotin ligase that converts biotin into a biotin-AMP complex that enables selective labeling of target proteins and pull down of it and proximal proteins
The step forward the paper makes is a more selective ligase, through directed evolution, and a split ligase as well. The split version of TurboID is composed of two fragments that are inactive by themselves and become active through protein–protein interactions or organelle–organelle interactions
The ability to map out proteomic interaction networks within a cell is important to understand the importance of spatial relationships within the cell
Genetics, genomics, and developmental biology
Heredity and variation.
Translation Initiation Site Profiling Reveals Widespread Synthesis of Non-AUG-Initiated Protein Isoforms in Yeast - http://cell.com/cell-systems/fulltext/S2405-4712(20)30240-4 - the Brar Lab at UC Berkeley discovered 149 genes whose expression begin at non-AUG start codons:
The paper used ribosome profiling, which enables in vivo identification of translated regions of the genome, to discover new open reading frames (ORF) in yeast. In combination with the profiling method, drugs that inhibit post-initiation ribosomes are used to identify initiation sites for translation.
These set of experiments generated large amounts of data that were annotated for ORFs with a focus on regions with non-AUG start sites - discovering 149 genes with protein isoforms that start at non-canonical sites
The utility and mechanism of these new sites are still outstanding questions, but the premise is that non-AUG start sides help cells respond to different conditions and use the same gene to generate more protein variants