Colorado is probably most well known for skiing, South Park, and the Buffs sports team. Under the radar, the University of Colorado, Boulder is home to some of the world’s greatest life scientists. With a world class cell therapy center mainly around the Gates Biomanufacturing Facility, one of the world’s best biochemists, and an interesting set of companies from Inscripta, DMC, and a lot more, Colorado is shaping to be an exciting place to start and build great businesses.

Cech Lab
Pioneering work on functional RNAs.Recent
Thomas Cech is an absolute legend - showing the catalytic nature of RNA, which validated the RNA world hypothesis. Recently, invented a method to screen and select for mutations that improve an RNA’s ability to be crystallized. Crystallography has been incredibly useful for proteins but for RNAs getting crystals that one can create diffraction maps with is difficult. To solve this problem, the Cech group designed a screening method whereby a pool of target RNAs with random mutations are crystallized and RNA amplification is used to match the best crystals with sequence - https://www.sciencedirect.com/science/article/pii/S0969212618301692 - the group did a proof-of-concept with a domain from a ribozyme in Tetrahymena - a valuable model organism given that its ribosomal RNAs are amplified ~10,000x more than other models. Ultimately, the ability to study the structures of various RNAs is important for general knowledge but the method of connecting structure to sequence for RNA and other nucleic acids is compelling to create new drug formats for catalysis, sensing, and delivery:

Conducted a study of single-cell variation in telomerase reverse transcriptase (TERT) expression - https://www.pnas.org/content/116/37/18488.short - TERT is an essential component of telomerase, which maintains telomere (ends of genome) length. The Cech group studied TERT expression across 10 human cancer lines finding a high level of variation in levels and localization potentially finding new avenues to treat cancer and increase lifespan. In general, single-cell tools are creating new data sets that help generate discoveries for new medicines.
Past
A short overview of the Cech Lab, starting in 1978 at Colorado working on catalytic RNAs in Tetrahymena - https://www.nature.com/articles/nrm.2017.105 - helping set the basis for the RNA origin of life.
A legendary paper showing the catalytic nature of RNA - http://dosequis.colorado.edu/Courses/MethodsLogic/papers/Cech1982.pdf
Led by Jamie Cate, determining the crystal structure of the catalytic core of Group I ribozymes, which enable self-splicing - https://science.sciencemag.org/content/273/5282/1678

With Joan Steitz, writing an outstanding review on noncoding RNAs - https://www.sciencedirect.com/science/article/pii/S0092867414003389 - particular on their various functions in the cell and the set of rules that govern utility across the diverse set - siRNAs, lncRNAs, and more:

Hoenger Lab
Using ET to study macromolecular machines.Recent
Profiled with the Lander Lab (Scripps), cryo-EM is an incredible tool to study molecular machines. On the other hand, the Hoenger Lab is pioneering the use of electron tomography to map out cellular compartments. The lab invented a metallothionein tag to enable high signal-to-noise detection of protein localization by EM - https://www.ncbi.nlm.nih.gov/pubmed/25974385
Using electron tomography (ET) to map out rabbit cardiomyocytes - https://www.sciencedirect.com/science/article/pii/S0079610716300372 - which are very dense and highly compartmentalized. Showing the power of ET in defining cellular structures from mitochondria to microtubules to other systems.
Past
Defining a protocol for cryo-ET, which is valuable to map 3D cell structures with high levels of variation - https://www.ncbi.nlm.nih.gov/pubmed/21842467
Copley Lab
Evolving enzymes for new functions.Recent
In E. coli, the Copley Lab studied the evolution of enzyme functions - https://www.biorxiv.org/content/10.1101/624205v1 - with a chassis that is defective in arginine synthesis and has an enzyme involved in proline synthesis with media supplemented with glucose and proline. After a few hundred generations, the enzyme involved in proline synthesis was amplified and diversified to help generate arginine as well. In the past, the lab did a similar Arg/Pro study - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2533218/ So instead of modifying enzymes in the arginine pathway, the bacteria duplicated a functioning enzyme and made it promiscuous enough to have a new function - make arginine:

Past
A good overview of the rules that drive enzyme function and diversification -http://www.jbc.org/content/287/1/3.long

Roop Lab
Driving the use of iPSCs for skin disease.Recent
Dennis Roop leads the Charles C. Gates Center for Regenerative Medicine and Stem Cell Biology (an interesting story of a rubber magnate with a passion for regenerative medicine), which has a gem hidden in it called the Gates Biomanufacturing Facility. Roop is actually at the Denver campus not Boulder, but his work and leadership in cell therapies is important to profile. The 14K square foot, GMP compliant facility is at the University of Colorado Anschutz Medical Campus. Established in 2015, the facility manufactures cell therapies and biologics for phase I/II clinical trials with a focus on:
Process development and documentation
Scaling
Manufacturing of both cell therapy- and protein-based therapeutics in compliance with cGMP standards
This top 5 facility (out of ~25 in the US) is a unique place for any company to partner with:

The Roop Lab did a recent review on the power of stem cell therapies to treat vitiligo and alopecia areata - https://www.jidonline.org/article/S0022-202X(19)31465-4/fulltext - in these indications and many autoimmune diseases melanocytes are often a target for damage. So various treatments like cell transplants, cell mobilization, and JAK inhibition (influence interferons) are interesting approaches to cure these diseases:

Past
A review of using induced pluripotent stem cells (iPSC) as models for dermatological diseases - https://www.ncbi.nlm.nih.gov/pubmed/25368014

Espinosa Lab
Functional genomics for disease.Recent
A useful review on cyclin dependent kinases (CDK), a field of expertise of the Espinosa Lab - https://www.ncbi.nlm.nih.gov/pubmed/30409083 - which are important to regulate transcription and could be interesting therapeutic targets:

With the setting that patients with Down syndrome are more susceptible to autoimmune disorders particular alopecia areata (AA; hair loss), the Espinosa Lab used the genetic connection between Trisomy 21 (Down Syndrome cause) and interferon activation to discover a JAK inhibitor, tofacitinib, has benefits for Down syndrome patients with AA - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6453941/
Past
Writing one of the canonical overviews on CDK8 - https://www.tandfonline.com/doi/full/10.4161/trns.1.1.12373 - a transcriptional activator influencing p53 and Wnt/β-catenin.
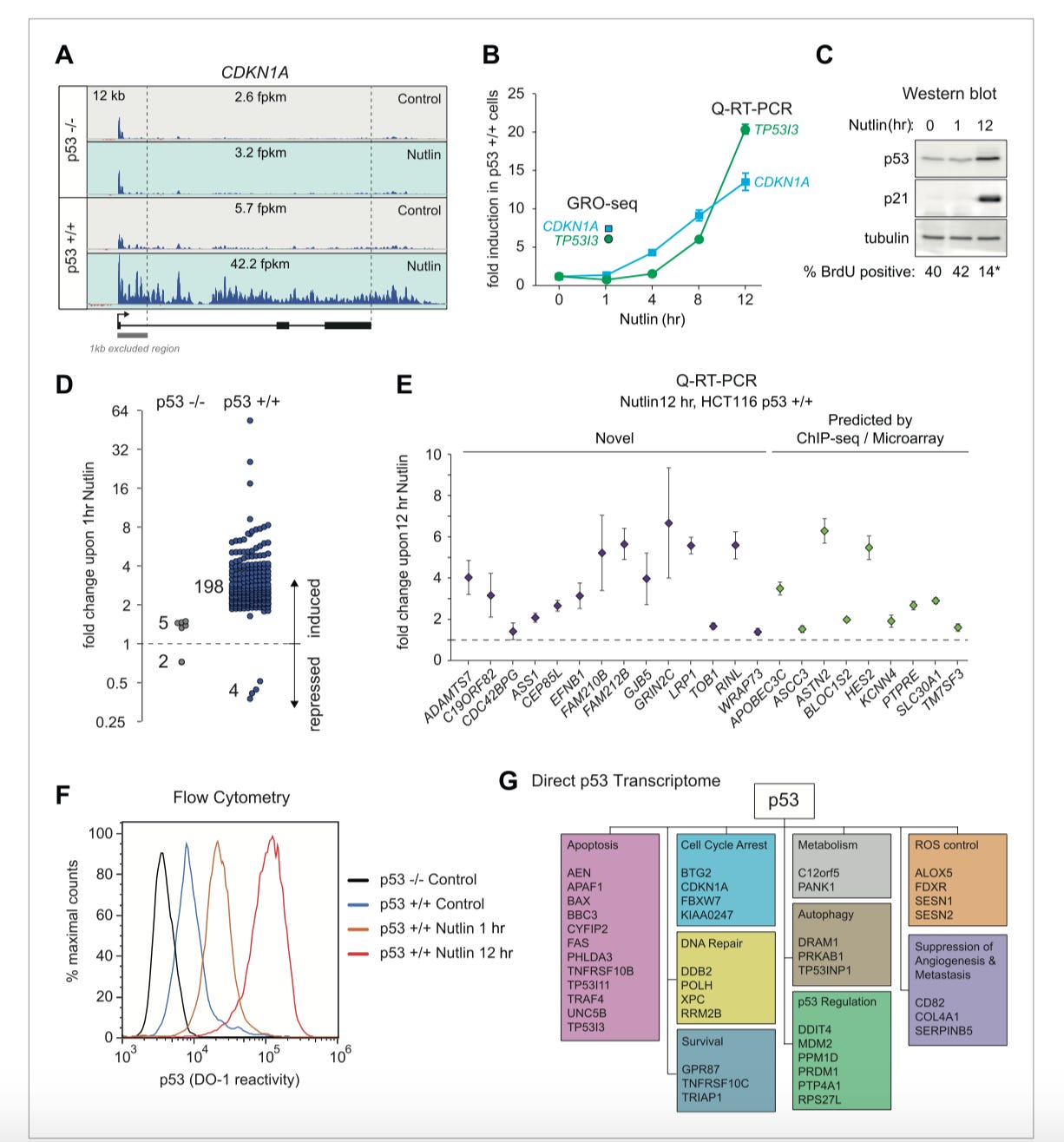
Invented a method called global run-on sequencing (GRO-seq) that measures transcription at the earliest stages instead of end products (i.e. microarrays) - https://cdn.elifesciences.org/articles/02200/elife-02200-v1.pdf - using it to do a genome-wide analysis of p53-regulated transcription where the tool can also be pointed toward other high value networks:

A fantastic review of p53 (one of the four horsemen of the undraggable genome) regulation of transcription and auto feedback from microRNAs and other RNA-binding proteins - https://academic.oup.com/bfg/article/12/1/46/186908



