Axial: https://linktr.ee/axialxyz
Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company. We are excited to be in business with you — email us at info@axialvc.com
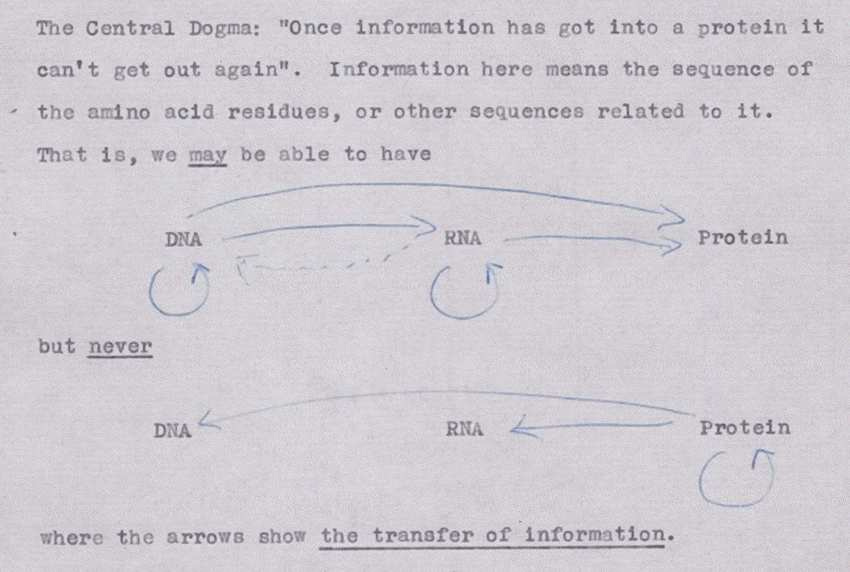
The central dogma of molecular biology has long been the foundation for our understanding of how biological information flows and is expressed. Originally postulated by Francis Crick, it defines the permissible directions of information transfer between the key biomolecules DNA, RNA, and proteins. However, as our knowledge has expanded, it has become clear that the central dogma provides an overly simplified view that overlooks crucial aspects of biological systems.
Sequence
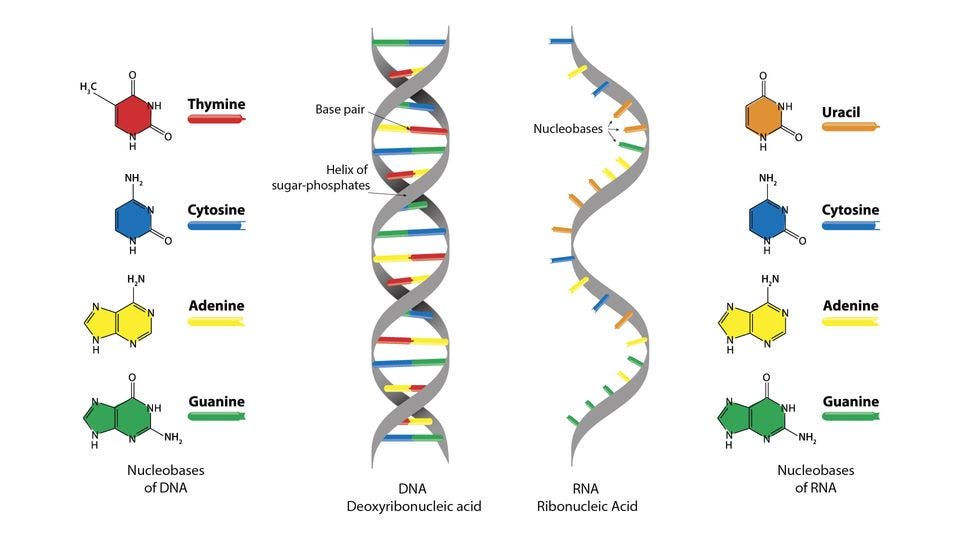
At the most fundamental level, biological information is encoded in the linear sequences of nucleotides in DNA and RNA, as well as the amino acid sequences of proteins. These sequences act as static blueprints, storing the core genetic instructions that determine an organism's inherited traits and the structure/function of biomolecules. The DNA sequence provides the master template that gets transcribed into messenger RNA, which is then translated by ribosomes into the corresponding sequence of amino acids to build functional proteins.
While sequences are relatively stable reservoirs of information, mutations can introduce variations that get propagated through successive generations. This is a driving force behind evolutionary processes, allowing organisms to adapt and diversify over long timescales. Additionally, certain mechanisms like RNA editing can alter sequences post-transcriptionally, adding another layer of regulation.
Structure
Moving beyond just the linear sequence, the next level of biological information lies in the precise three-dimensional structures adopted by biomolecules like proteins and nucleic acids. The structure of a protein largely determines its function by defining features like catalytic sites, binding pockets, and interactions with other biomolecules.
In contrast to the static nature of sequences, biomolecular structures exhibit semi-dynamic properties. While folding into specific structural conformations based on their sequences, proteins and nucleic acids can undergo induced conformational changes that alter their activities and binding properties. These structural rearrangements are often regulated by cellular signals or environmental cues, enabling biological responsiveness.
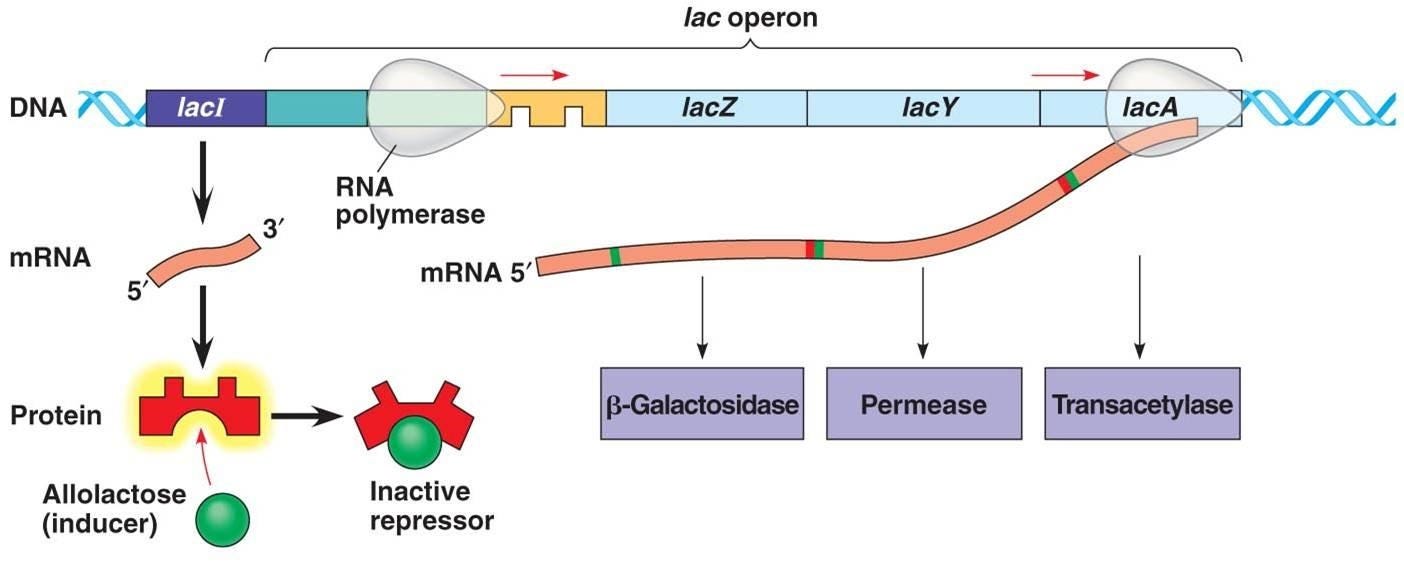
A key example highlighted is the lac operon system in bacteria, where the lac repressor protein changes its structure upon binding to lactose, releasing its grip on the DNA to allow gene transcription. Such allostery and induced fit mechanisms are fundamental to many regulatory processes in biology.
Systems
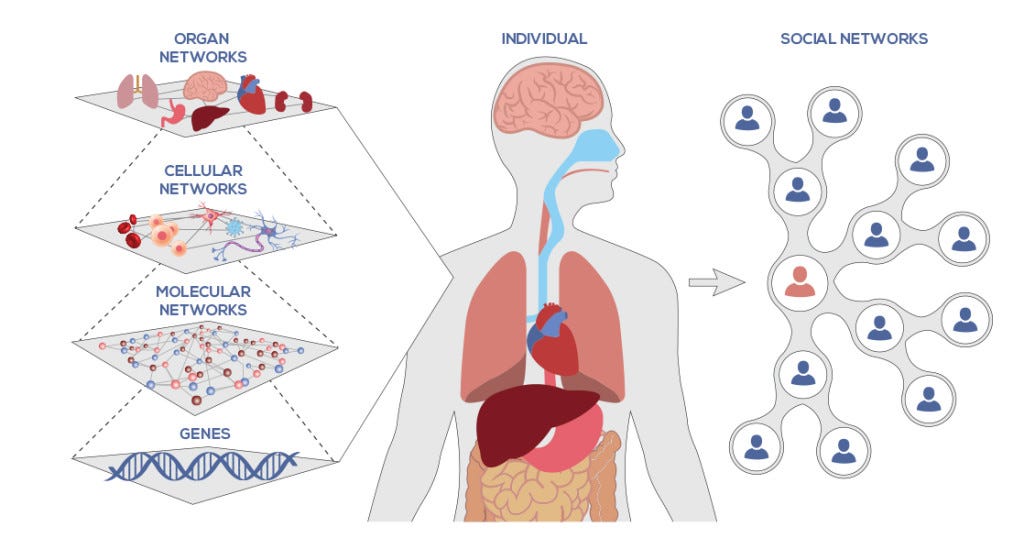
The third and most dynamic modality of biological information encompasses the concentrations and abundances of different biomolecules present within cells and organisms at any given time. This "systems" level captures the molecular composition and concentrations of proteins, metabolites, RNAs and other biomolecules - which are in constant flux based on gene expression, enzyme activities, and regulatory feedback loops.
In contrast to the relatively fixed sequences and semi-flexible structures, these concentration levels are highly dynamic and responsive to both intracellular requirements and environmental perturbations. Gene regulatory networks modulate the transcription of genes based on cellular signals, adjusting protein levels to meet metabolic demands. Metabolic pathways are regulated by the concentrations of enzymes, substrates and products in a continuous dance of feedback loops.
The lac operon example again illustrates this principle, wherein the expression of the lac genes (and thus the levels of the encoded proteins) is regulated based on the relative abundances of glucose and lactose in the environment. Only when glucose is depleted and lactose is available does the system upregulate the machinery to metabolize lactose as an alternative carbon source.
While presented as distinct modalities, sequence, structure, and systems are inherently interconnected in facilitating the intricate molecular choreography within cells. DNA sequences encode the amino acid sequences of proteins, which fold into specific structures that determine their functions and interactions. These proteins, in turn, act as catalysts, regulators, and components of cellular systems - with their abundances and activities modulated by various regulatory mechanisms. Modeling biological systems necessitates approximating how these different modalities dynamically interact to manifest an organism's phenotype or observable characteristics. Current experimental techniques allow accurate measurements of sequences, structures, and abundance levels. However, the true challenge lies in capturing and simulating the intervening transitions between these measured states over time and in response to environmental inputs.
Biological systems operate far from equilibrium, with intricate networks of regulatory circuits and feedback loops continuously adjusting protein levels, activating or repressing genes, and modifying metabolic fluxes. Even subtle perturbations can propagate through these highly interconnected systems, producing cascading effects that shape the emergent behavior.
Two critical variables highlighted are time and the environment, which are often overlooked in oversimplified models of biological information flow. Living systems are inherently dynamic, functioning within the dimension of time rather than as a static snapshot. Biological processes like development, circadian rhythms, and cellular responses all unfold over varying timescales. Moreover, these processes do not occur in isolation but are profoundly influenced by external environmental factors as well as the organism's own actions. Environmental cues like temperature, nutrient availability, or signaling molecules can trigger regulatory cascades that reprogram gene expression, metabolic activity, and other systemic adjustments. Conversely, organisms can actively modify and shape their environment through processes like nutrient foraging, niche construction, or anthropogenic activities on a global scale.
This bidirectional exchange and feedback between an organism and its environment is a hallmark of biological systems, adding considerable complexity to efforts aimed at comprehensive modeling and prediction.
The concepts of sequence, structure, and systems provide a useful framework for conceptualizing the multi-layered nature of biological information and how it is processed across different scales. However, biology operates through a highly integrated and dynamic interplay between these modalities, unfolding over time and intimately coupled with environmental context.
Reductionist approaches that focus solely on mapping sequences to structures or abundances are likely to fall short in capturing the full richness and complexity of living systems. A more holistic perspective is needed, one that embraces the inherent non-linearity, feedback loops, and multi-scale interactions that give rise to the astounding diversity and adaptability of life on Earth.
Interdisciplinary efforts bridging biology, physics, computer science, and information theory may provide avenues for developing integrative models and computational frameworks better suited for grappling with these challenges. As our ability to collect multi-dimensional, time-resolved data improves, perhaps coupled with advanced machine learning and simulation techniques, we may inch closer to reconstructing and predicting how biological information flows and manifests across sequences, structures, and systems.
Ultimately, deepening our understanding of these fundamental principles could pave the way for transformative applications, from rationally designing biomolecules and pathways to engineering robust biological systems resilient to environmental perturbations - endeavors crucial for addressing pressing challenges in healthcare, agriculture, energy, and environmental sustainability.