Rational design of antibodies targeting specific epitopes within intrinsically disordered proteins
Inventors & their inventions
Axial: https://linktr.ee/axialxyz
Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company. We are excited to be in business with you — email us at info@axialvc.com
This work presents an innovative computational method for rationally designing antibodies to target specific epitopes within intrinsically disordered proteins (IDPs) and disordered regions of proteins. IDPs lack stable tertiary structure and exist as heterogeneous ensembles of conformations. This dynamic and disordered nature makes generating antibodies against specific epitopes within IDPs very challenging using conventional techniques like immunization. The authors' rational design strategy overcomes these challenges by enabling the precise targeting of desired epitopes in IDPs.
A key idea behind the design method is to identify short peptide sequences complementary to the target epitope that can enforce a β-strand conformation on the epitope upon binding. These complementary peptides are then grafted into the complementarity determining regions (CDRs) of an antibody scaffold to generate antibodies designed to bind the target epitope.
The process for identifying complementary peptides involves analyzing known protein structures to find fragments that form β-strand interactions with subsequences matching the target epitope. These fragments are then combined using a "cascade method" to build full-length complementary peptides matching the length of the target epitope, while ensuring the proper parallel or anti-parallel β-strand hydrogen bonding pattern is maintained. The authors demonstrate the generality of this approach by analyzing its coverage of all possible 8-residue epitopes within the IDP α-synuclein and the amyloidogenic peptides Aβ42 and IAPP. They find that for these disordered systems, over 95% of residue positions are covered by at least one predicted complementary peptide, with a median of 200 peptides per position. This indicates the method can generate multiple complementary peptide candidates for virtually any desired epitope in these IDPs.
An advantage is that the complementary peptides are derived from actual β-strand interfaces observed in structured proteins in the Protein Data Bank. This biological relevance increases the likelihood that the designed complementary peptides can form the intended β-strand interactions with the target epitope. The authors further evaluate cross-reactivity by searching the complementary peptides against the human proteome, finding only 0.2% directly match human sequences, suggesting high specificity.
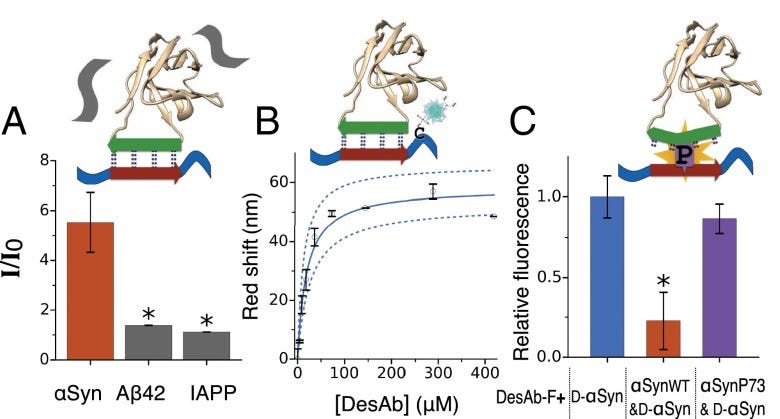
With complementary peptides designed for α-synuclein, Aβ42, and IAPP, the authors graft these into the CDR3 loop of a stable human single-domain antibody scaffold. Biophysical characterization (CD, SDS-PAGE) confirms the structural integrity of the grafted antibody designs, which are termed "DesAbs" (Designed Antibodies). The key experimental results validate that the DesAbs bind their intended targets with good specificity, as demonstrated by ELISA binding assays and a dot blot analysis comparing binding to cell lysates with and without the target antigen. For the α-synuclein targeted DesAb-F, a fluorescence competition assay provides strong evidence that binding specifically occurs at the designed epitope, as an α-synuclein mutant with a proline insertion in the epitope abolishes binding.
Importantly, the authors show the DesAb-F can potently inhibit α-synuclein aggregation, even at substoichiometric concentrations relative to the protein. Seeded aggregation assays reveal a concentration-dependent inhibition of the elongation phase by DesAb-F. This highlights a key advantage of targeting specific epitopes in aggregation-prone IDPs - the designed antibodies can bind and cap elongating fibril ends to inhibit further growth. While the affinity of the single CDR-grafted DesAb-F for monomeric α-synuclein (Kd ~20 μM) is moderate, this may actually be ideal for inhibiting aggregation versus simply sequestering monomers. To illustrate how affinity can be substantially improved, the authors design a "two-loop" DesAb by grafting complementary peptides into both the CDR2 and CDR3 loops of the scaffold, thereby increasing the interaction interface. This dual-grafted antibody exhibits picomolar affinity for α-synuclein, comparable to traditional antibodies.
Overall, this rational design strategy enables the precise targeting of antibodies against epitopes within IDPs that would be extremely challenging using conventional techniques. The ability to design antibodies against any disordered epitope of interest has immense potential for both research applications to probe IDP structure/function, as well as therapeutic applications targeting disease-associated IDPs.
Some key strengths of the method include:
1) Generality and coverage - The computational analysis shows the approach can target nearly any epitope within the IDPs studied, with multiple complementary peptide options per epitope.
2) Biological relevance - Deriving complementary peptides from known β-strand interfaces increases the likelihood of productive binding to the disordered target.
3) Specificity - The human proteome analysis suggests the designed complementary peptides are highly specific for the intended target.
4) Modulation of activity - The α-synuclein DesAb can potently inhibit aggregation, illustrating how the designs can potentially modulate pathological activities of IDPs.
5) Tailoring affinity - Grafting complementary peptides into multiple CDR loops enables tuning of the antibody affinity over a wide range as desired.
Some potential limitations and areas for further development include stability of multi-loop grafts. The two-loop DesAb exhibited reduced stability compared to the single-loop version, suggesting multi-loop designs may require more stable antibody scaffolds. Also, the moderate affinities of the single-loop DesAbs precluded Western blot detection, though the higher affinity two-loop version was successful. While the in vitro characterization is very promising, evaluating the designs in more physiologically relevant cellular contexts will be an important next step. And Further affinity maturation; the authors note that combining the rational design approach with directed evolution techniques like phage display could enable further affinity improvements. While IDPs were the focus, the strategy could potentially extend to targeting specific epitopes in ordered proteins as well.
Ultimately, this paper makes an important conceptual and technical advance by introducing a rational, structure-based computational strategy to design antibodies precisely targeting desired epitopes within disordered proteins. By grafting complementary peptides predicted to bind the epitope in a β-strand conformation, the authors demonstrate successful design of antibodies against three different IDPs associated with neurodegenerative diseases. The approach opens up exciting new avenues for generating antibodies to probe biological roles of IDPs and potentially develop therapeutics targeting disease-linked disordered proteins and epitopes. While some remaining challenges exist, this design strategy creates powerful new opportunities in the realm of antibody engineering and drug design.