Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage
Surveying great inventors and businesses
Axial: https://linktr.ee/axialxyz
Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company. We are excited to be in business with you — email us at info@axialvc.com
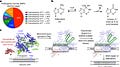
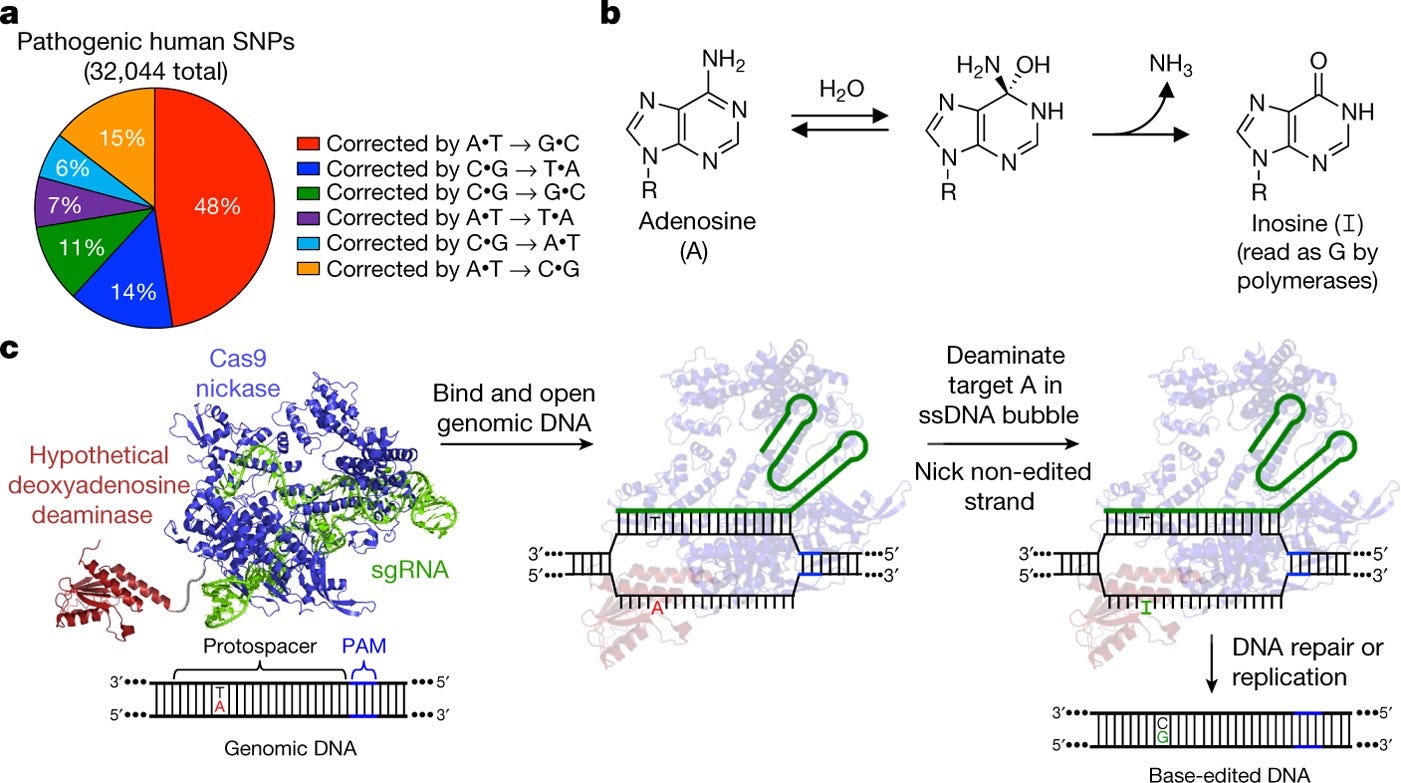
This groundbreaking paper describes the development of a new class of genome editing proteins called adenine base editors (ABEs) that can directly convert A•T base pairs to G•C base pairs in genomic DNA without making double-stranded DNA breaks. This greatly expands the capabilities of base editing technology, which previously could only convert C•G to T•A.
The authors started with the E. coli tRNA adenosine deaminase enzyme TadA, which naturally deaminates adenosine to inosine in tRNA molecules. However, TadA has no activity on DNA. Through seven rounds of protein engineering efforts using a bacterial selection system, they were able to evolve TadA variants that could deaminate adenosines in DNA when fused to a catalytically inactive Cas9 nickase.
The key steps in evolving highly active and specific ABEs were:
1) Identifying initial TadA mutations like A106V and D108N that enabled some low-level DNA deaminase activity (ABE1).
2) Introducing additional mutations like D147Y and E155V to improve activity (ABE2) as well as mutations to dimerize TadA (ABE2.9).
3) Further accumulating mutations like L84F, H123Y, I157F that increased activity up to 29% editing at some sites (ABE3.1).
4) Mutating residues that interact with DNA like A142N to broaden the targetable sequence motifs from just YAC to other NAN contexts (ABE4.3).
5) Switching to a heterodimer with wildtype TadA providing a structural role (ABE5.3) to avoid accumulation of too many mutations impairing complex formation.
6) Using DNA shuffling (ABE6.3) and identifying mutations like P48S that further improved activity up to 47%.
7) Finally evolving additional mutations like W23R, R152H that together enabled very high efficiencies averaging 53-58% editing across a wide variety of NAN sequence contexts (ABE7.10).
The evolved ABE7.10 could edit target A•T base pairs in human genomic DNA with 50% or higher efficiency at 11 out of 17 tested sites, with very high product purity (typically >99.9%) and extremely low rates of indels or unwanted byproduct edits (<0.1%). This compares favorably to the efficiencies and purity of current cytosine base editors like BE3.
Importantly, ABE7.10 outperformed a current Cas9 nuclease-based homology-directed repair approach by over 1000-fold in terms of the ratio of desired edits to unwanted indel byproducts. Unlike BE3 which requires strong inhibition of DNA repair to prevent excision of uracil intermediates, the authors found that the inosine intermediates generated by ABEs are inefficiently processed, obviating the need for inhibition strategies.
Analysis of DNA sequencing reads indicated that while early-stage ABEs edited nearby adenines relatively independently, the late-stage ABEs like ABE7.10 tended to edit nearby adenines more processively within their 4-6 nt editing window. This processivity likely contributes to their high product purity.
Interestingly, when tested at known off-target sites identified for their guide RNAs from GUIDE-seq data, the authors found that ABE7 variants induced detectable off-target edits at only 33% of these sites, with an average efficiency of just 1.3%. In contrast, Cas9 nuclease induced detectable indels at 75% of these sites with 14% average efficiency. This suggests ABEs may induce lower off-target effects than Cas9, although unbiased profiling methods will be needed to quantify their full specificity profiles.
As a key demonstration, the authors used ABE7.10 to install two different types of genetic modifications relevant to human disease. First, they were able to introduce mutations in the promoters of the HBG1 and HBG2 genes with about 30% efficiency that are known to cause hereditary persistence of fetal hemoglobin, a condition that can ameliorate sickle cell disease and beta-thalassemia. Second, in patient-derived cells harboring the C282Y mutation in HFE that causes hereditary hemochromatosis, they were able to correct this point mutation back to wildtype with 28% efficiency and high purity.