Blueprinting extendable nanomaterials with standardized protein blocks
Inventors & their inventions
Axial: https://linktr.ee/axialxyz
Axial partners with great founders and inventors. We invest in early-stage life sciences companies such as Appia Bio, Seranova Bio, Delix Therapeutics, Simcha Therapeutics, among others often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company. We are excited to be in business with you — email us at info@axialvc.com
This groundbreaking paper describes a major advance in the field of protein design and nanotechnology, introducing a novel platform for constructing modular protein nanomaterials with programmable size and geometry. The key innovation is the development of "twistless helix repeat" (THR) protein building blocks that conform to simple geometrical standards, enabling the rational design of complex protein assemblies much like building a house from uniform lumber pieces.
Prior work on designing protein nanomaterials has been hindered by the irregular, complex shapes of natural proteins, requiring computationally intensive sampling to identify complementary interfaces for each design case. The assemblies generated have generally been limited in size and cannot be readily scaled up or down. In contrast, the THR platform achieves remarkable simplicity by restricting the design to ideal, straight α-helices aligned in parallel, avoiding the irregularities inherent to most protein structures.
The THR building blocks consist of repeating units constructed from identical straight α-helices, with modular linear, curved, and angled geometries specified by varying three key parameters: the distance between helix axes, the vertical step between repeats, and the angle of rotation between repeats. Remarkably, changing just the rotation angle transforms the building block from a linear to a curved trajectory, generating arcs with different radii of curvature depending on the angle value. Additionally, "turn modules" introduce abrupt changes in direction by incorporating helices at different phase angles relative to the preceding helix, enabling construction of polygonal and circular shapes.
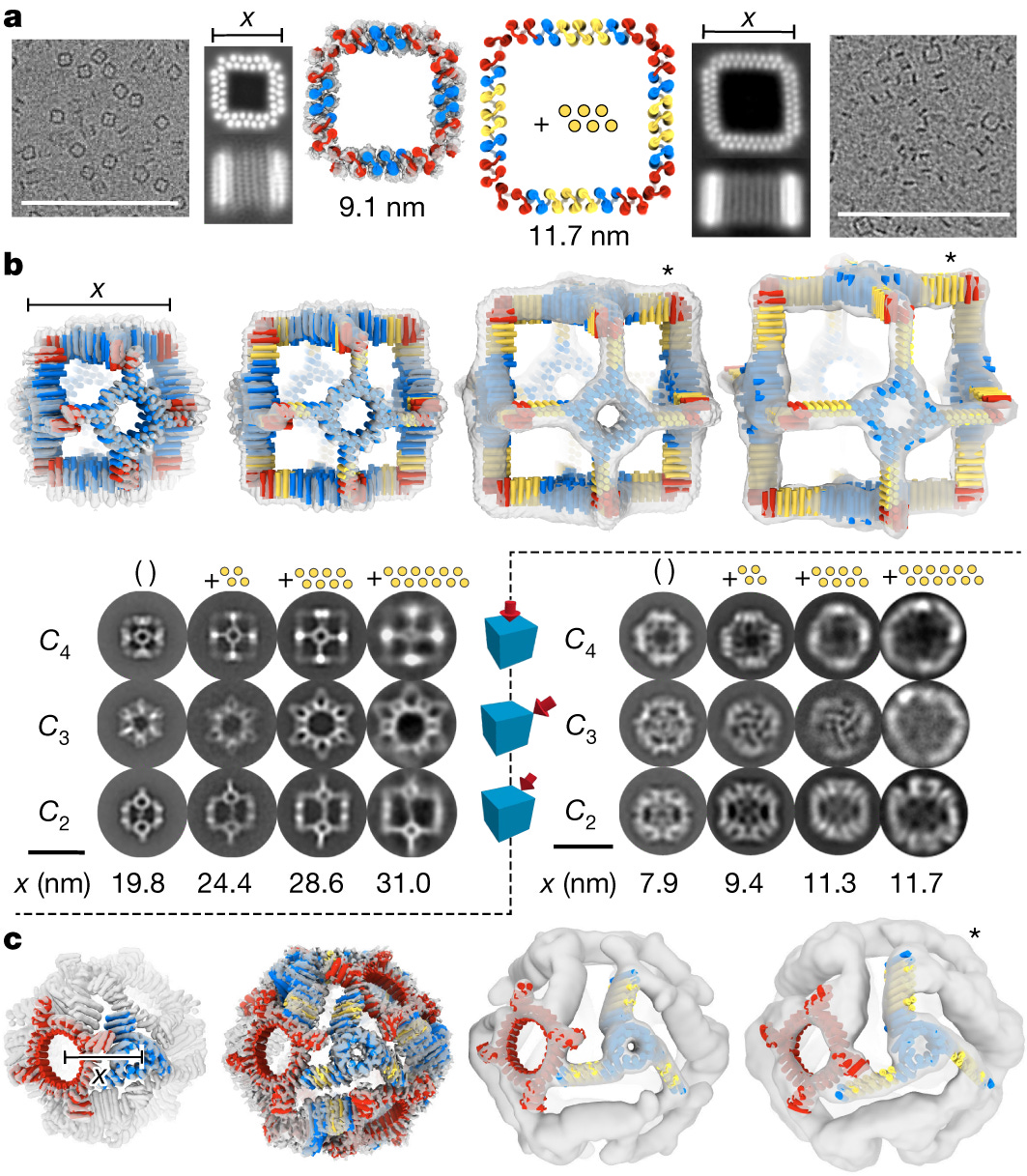
A key strength of this platform is the ability to incorporate standardized interaction interfaces between building blocks while maintaining the simple geometric forms, achieved through careful sequence design. Specific examples demonstrated include polygonal and circular oligomers formed by self-assembly of turn modules, as well as larger polyhedral cage architectures assembled from circular "ring" building blocks connected by linear "arm" blocks at defined angles dictated by the polyhedral symmetry. The success of these designs, as validated by cryogenic electron microscopy (cryo-EM) and X-ray crystallography data, is a testament to the power of this rational, blueprint-like approach compared to traditional computational protein-protein interface design methods.
Perhaps the most remarkable feature of the THR platform, however, is its ability to produce nanomaterials with readily adjustable, expandable dimensions simply by varying the number of repeating units in the constituent building blocks. This is a stunning achievement, as it overcomes a major limitation of previous protein design methods which generated assemblies with fixed, pre-determined sizes. The modularity and idealized helical geometries of the THR blocks enable resizing of designed materials along one, two or three dimensions with minimal structural perturbation, akin to a construction worker lengthening a wall by adding more uniform bricks.
The paper provides impressive experimental evidence for the scalability of THR nanomaterials across multiple architectures. Polygonal assemblies could be expanded by inserting additional repeat units, retaining the designed angles but with clearly enlarged edge lengths. For polyhedral cage structures, both the THR "arm" extensions and the rings forming the cage faces could be linearly extended, resulting in a series of cages with a perfectly preserved overall cubic shape but incrementally larger volumes, as confirmed by cryo-EM reconstructions. Remarkably, even complex two-component cage architectures consisting of different oligomeric rings interconnected by THR arms could be resized in a predictable manner. The ability to increase dimensions while maintaining the precise geometric blueprint is an exceptional accomplishment with tremendous potential for creating tailored nanomaterials and investigating size-dependent physical properties.
Moreover, the authors demonstrate the capability to reinforce the designed structures by incorporating rigid "buttressing" elements that bridge different components, maintaining the overall architecture while increasing stability. Concentric ring assemblies were constructed by fusing different sized rings with a linear spacer scaffold. For polyhedral cages, cryo-EM data confirmed that the idealized junctions between rings and THR arms resulted in the requisite inter-component angles for perfect polyhedral assembly, even in the absence of traditional "knob-into-hole" interface interactions.
While the THR platform represents a paradigm shift for protein nanotechnology, there are some limitations to acknowledge. The requirement for parallel, straight α-helices imposes geometric constraints that may not be suitable for every desired architecture. However, the authors propose promising strategies to overcome such limitations, such as incorporating heterodimeric or heterotrimeric interfaces between modules to break symmetry and increase addressability. Additionally, the high-symmetry assemblies demonstrated could potentially suffer from defects or incomplete formation as complexity increases, an issue that may necessitate solid-phase synthesis approaches to improve product homogeneity.
Looking ahead, the THR platform opens exciting avenues for creating functional protein-based nanomaterials by design. The ability to readily adjust dimensions enables systematic investigation of size-dependent properties, while the modular nature allows incorporation of diverse functional domains arranged with atomic precision. Applications could span drug delivery, sensing, catalysis, molecular electronics, and more. The blueprinting approach may even facilitate integration with other nanomaterial systems like DNA origami through interface design.
Moreover, the fundamental insights into programming protein structural geometry have implications beyond nanotechnology. Certain protein misfolding diseases are linked to improper twisting of amyloidogenic proteins; the principles for enforcing linear, untwisted helical trajectories could lead to stabilized versions resistant to pathological conformations. From a basic science perspective, the ability to construct such geometrically idealized protein architectures provides a powerful platform for exploring how sequence encodes three-dimensional structure and assembly.
This groundbreaking study represents a tour de force of computational protein design leveraging physical principles to achieve unprecedented control over material architecture. The development of standardized, extendable protein building blocks that can be rationally assembled into diverse geometries of controllable size is a gamechanger for the field of protein nanotechnology. With a versatile toolkit now available for blueprinting precise nanomaterials from polypeptide chains, one can envision virtually unlimited potential for creating functional, tailored biomolecular machines by design. Overcoming the "complexity and lack of symmetry" that once defined protein structure, this pioneering work establishes a powerful new paradigm in structural design at the nanoscale.